Generating Synthetic Data Sets with ‘synthpop’ in R
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.

Synthpop – A great music genre and an aptly named R package for synthesising population data. I recently came across this package while looking for an easy way to synthesise unit record data sets for public release. The goal is to generate a data set which contains no real units, therefore safe for public release and retains the structure of the data. From which, any inference returns the same conclusion as the original. This will be a quick look into synthesising data, some challenges that can arise from common data structures and some things to watch out for.

Sythesising data
This example will use the same data set as in the synthpop documentation and will cover similar ground, but perhaps an abridged version with a few other things that weren’t mentioned. The SD2011 contains 5000 observations and 35 variables on social characteristics of Poland. A subset of 12 of these variables are considered.
suppressPackageStartupMessages(library(synthpop))
suppressPackageStartupMessages(library(tidyverse))
suppressPackageStartupMessages(library(sampling))
suppressPackageStartupMessages(library(partykit))
mycols <- c("darkmagenta", "turquoise")
options(xtable.floating = FALSE)
options(xtable.timestamp = "")
myseed <- 20190110
# filtering the dataset
original.df <- SD2011 %>% dplyr::select(sex, age, socprof, income, marital, depress, sport, nofriend, smoke, nociga, alcabuse, bmi)
head(original.df)
## sex age socprof income marital depress sport nofriend smoke
## 1 FEMALE 57 RETIRED 800 MARRIED 6 NO 6 NO
## 2 MALE 20 PUPIL OR STUDENT 350 SINGLE 0 NO 4 NO
## 3 FEMALE 18 PUPIL OR STUDENT NA SINGLE 0 NO 20 NO
## 4 FEMALE 78 RETIRED 900 WIDOWED 16 YES 0 NO
## 5 FEMALE 54 SELF-EMPLOYED 1500 MARRIED 4 YES 6 YES
## 6 MALE 20 PUPIL OR STUDENT -8 SINGLE 5 NO 10 NO
## nociga alcabuse bmi
## 1 -8 NO 30.79585
## 2 -8 NO 23.44934
## 3 -8 NO 18.36547
## 4 -8 NO 30.46875
## 5 20 NO 20.02884
## 6 -8 NO 23.87511
The objective of synthesising data is to generate a data set which resembles the original as closely as possible, warts and all, meaning also preserving the missing value structure. There are two ways to deal with missing values 1) impute/treat missing values before synthesis 2) synthesise the missing values and deal with the missings later. The second option is generally better since the purpose the data is supporting may influence how the missing values are treated.
Missing values can be simply NA or some numeric code specified by the collection. A useful inclusion is the syn function allows for different NA types, for example income, nofriend and nociga features -8 as a missing value. A list is passed to the function in the following form.
# setting continuous variable NA list cont.na.list <- list(income = c(NA, -8), nofriend = c(NA, -8), nociga = c(NA, -8))
By not including this the -8’s will be treated as a numeric value and may distort the synthesis.
After synthesis, there is often a need to post process the data to ensure it is logically consistent. For example, anyone who is married must be over 18 and anyone who doesn’t smoke shouldn’t have a value recorded for ‘number of cigarettes consumed’. These rules can be applied during synthesis rather than needing adhoc post processing.
# apply rules to ensure consistency
rules.list <- list(
marital = "age < 18",
nociga = "smoke == 'NO'")
rules.value.list <- list(
marital = "SINGLE",
nociga = -8)
The variables in the condition need to be synthesised before applying the rule otherwise the function will throw an error. In this case age should be synthesised before marital and smoke should be synthesised before nociga.
There is one person with a bmi of 450.
SD2011[which.max(SD2011$bmi),] ## sex age agegr placesize region edu ## 3953 FEMALE 58 45-59 URBAN 20,000-100,000 Lubelskie SECONDARY ## eduspec socprof unempdur income ## 3953 economy and administration LONG-TERM SICK/DISABLED -8 1300 ## marital mmarr ymarr msepdiv ysepdiv ls depress ## 3953 MARRIED 4 1982 NA NA MOSTLY SATISFIED 1 ## trust trustfam trustneigh sport nofriend smoke ## 3953 ONE CAN`T BE TOO CAREFUL YES YES YES 2 NO ## nociga alcabuse alcsol workab wkabdur wkabint wkabintdur emcc englang ## 3953 -8 NO NO NO -8 NO <NA> <NA> NONE ## height weight bmi ## 3953 149 NA 449.9797
Their weight is missing from the data set and would need to be ![]() for this to be accurate. I don’t believe this is correct! So, any bmi over 75 (which is still very high) will be considered a missing value and corrected before synthesis.
for this to be accurate. I don’t believe this is correct! So, any bmi over 75 (which is still very high) will be considered a missing value and corrected before synthesis.
# getting around the error: synthesis needs to occur before the rules are applied original.df$bmi <- ifelse(original.df$bmi > 75, NA, original.df$bmi)
Consider a data set with ![]() variables. In a nutshell, synthesis follows these steps:
variables. In a nutshell, synthesis follows these steps:
- Take a simple random sample of
 and set as
and set as 
- Fit model
 and draw
and draw  from
from 
- Fit model
 and draw
and draw  from
from 
- And so on, until

The data can now be synthesised using the following code.
# synthesise data synth.obj <- syn(original.df, cont.na = cont.na.list, rules = rules.list, rvalues = rules.value.list, seed = myseed) ## ## Unexpected values (not obeying the rules) found for variable(s): nociga. ## Rules have been applied but make sure they are correct. ## Synthesis ## ----------- ## sex age socprof income marital depress sport nofriend smoke nociga ## alcabuse bmi synth.obj ## Call: ## ($call) syn(data = original.df, rules = rules.list, rvalues = rules.value.list, ## cont.na = cont.na.list, seed = myseed) ## ## Number of synthesised data sets: ## ($m) 1 ## ## First rows of synthesised data set: ## ($syn) ## sex age socprof income marital depress sport ## 1 FEMALE 45 EMPLOYED IN PUBLIC SECTOR 1500 SINGLE 5 YES ## 2 MALE 65 OTHER ECONOMICALLY INACTIVE 500 SINGLE 5 YES ## 3 FEMALE 17 PUPIL OR STUDENT NA SINGLE 3 NO ## 4 FEMALE 48 EMPLOYED IN PRIVATE SECTOR 1000 MARRIED 0 NO ## 5 MALE 50 EMPLOYED IN PRIVATE SECTOR 1300 MARRIED 0 NO ## 6 FEMALE 65 RETIRED 1200 WIDOWED 6 NO ## nofriend smoke nociga alcabuse bmi ## 1 3 NO -8 NO 26.12245 ## 2 30 NO -8 NO 29.32099 ## 3 7 NO -8 NO 22.40588 ## 4 10 NO -8 NO 25.81663 ## 5 12 YES 20 YES 27.17063 ## 6 15 NO -8 NO 27.51338 ## ... ## ## Synthesising methods: ## ($method) ## sex age socprof income marital depress sport nofriend ## "sample" "cart" "cart" "cart" "cart" "cart" "cart" "cart" ## smoke nociga alcabuse bmi ## "cart" "cart" "cart" "cart" ## ## Order of synthesis: ## ($visit.sequence) ## sex age socprof income marital depress sport nofriend ## 1 2 3 4 5 6 7 8 ## smoke nociga alcabuse bmi ## 9 10 11 12 ## ## Matrix of predictors: ## ($predictor.matrix) ## sex age socprof income marital depress sport nofriend smoke ## sex 0 0 0 0 0 0 0 0 0 ## age 1 0 0 0 0 0 0 0 0 ## socprof 1 1 0 0 0 0 0 0 0 ## income 1 1 1 0 0 0 0 0 0 ## marital 1 1 1 1 0 0 0 0 0 ## depress 1 1 1 1 1 0 0 0 0 ## sport 1 1 1 1 1 1 0 0 0 ## nofriend 1 1 1 1 1 1 1 0 0 ## smoke 1 1 1 1 1 1 1 1 0 ## nociga 1 1 1 1 1 1 1 1 1 ## alcabuse 1 1 1 1 1 1 1 1 1 ## bmi 1 1 1 1 1 1 1 1 1 ## nociga alcabuse bmi ## sex 0 0 0 ## age 0 0 0 ## socprof 0 0 0 ## income 0 0 0 ## marital 0 0 0 ## depress 0 0 0 ## sport 0 0 0 ## nofriend 0 0 0 ## smoke 0 0 0 ## nociga 0 0 0 ## alcabuse 1 0 0 ## bmi 1 1 0
The compare function allows for easy checking of the sythesised data.
# compare the synthetic and original data frames compare(synth.obj, original.df, nrow = 3, ncol = 4, cols = mycols)$plot

Solid. The distributions are very well preserved. Did the rules work on the smoking variable?
# checking rules worked
table(synth.obj$syn[,c("smoke", "nociga")])
## nociga
## smoke -8 1 2 3 4 5 6 7 8 10 12 14 15
## YES 13 13 13 32 5 49 17 12 28 308 26 3 135
## NO 3777 0 0 0 0 0 0 0 0 0 0 0 0
## nociga
## smoke 16 18 20 21 22 24 25 26 29 30 35 40 50
## YES 5 7 418 2 1 2 31 2 2 51 1 33 1
## NO 0 0 0 0 0 0 0 0 0 0 0 0 0
## nociga
## smoke 60
## YES 1
## NO 0
They did. All non-smokers have missing values for the number of cigarettes consumed.
compare can also be used for model output checking. A logistic regression model will be fit to find the important predictors of depression. The depression variable ranges from 0-21. This will be converted to
- 0-7 – no evidence of depression (0)
- 8-21 – evidence of depression (1)
This split leaves 3822 (0)’s and 1089 (1)’s for modelling.
# ------------ MODEL COMPARISON glm1 <- glm.synds(ifelse(depress > 7, 1, 0) ~ sex + age + log(income) + sport + nofriend + smoke + alcabuse + bmi, data = synth.obj, family = "binomial") ## Warning in log(income): NaNs produced summary(glm1) ## Warning: Note that all these results depend on the synthesis model being correct. ## ## Fit to synthetic data set with a single synthesis. ## Inference to coefficients and standard errors that ## would be obtained from the observed data. ## ## Call: ## glm.synds(formula = ifelse(depress > 7, 1, 0) ~ sex + age + log(income) + ## sport + nofriend + smoke + alcabuse + bmi, family = "binomial", ## data = synth.obj) ## ## Combined estimates: ## xpct(Beta) xpct(se.Beta) xpct(z) Pr(>|xpct(z)|) ## (Intercept) -1.00819811 0.71605764 -1.4080 0.1591356 ## sexFEMALE 0.35681507 0.10010909 3.5643 0.0003649 *** ## age 0.09004527 0.00384758 23.4031 < 2.2e-16 *** ## log(income) -0.68041355 0.08829602 -7.7060 1.298e-14 *** ## sportNO -0.66446597 0.11880451 -5.5929 2.233e-08 *** ## nofriend 0.00028325 0.00679104 0.0417 0.9667305 ## smokeNO 0.08544511 0.11894243 0.7184 0.4725269 ## alcabuseNO -0.72124437 0.21444108 -3.3634 0.0007700 *** ## bmi 0.00644972 0.01036016 0.6226 0.5335801 ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 # compare to the original data set compare(glm1, original.df, lcol = mycols) ## Warning in log(income): NaNs produced ## ## Call used to fit models to the data: ## glm.synds(formula = ifelse(depress > 7, 1, 0) ~ sex + age + log(income) + ## sport + nofriend + smoke + alcabuse + bmi, family = "binomial", ## data = synth.obj) ## ## Differences between results based on synthetic and observed data: ## Std. coef diff p value CI overlap ## (Intercept) -1.26517500 0.206 0.6772453 ## sexFEMALE 0.27373709 0.784 0.9301678 ## age 0.85530291 0.392 0.7818065 ## log(income) 0.98568572 0.324 0.7485449 ## sportNO 0.16485706 0.869 0.9579439 ## nofriend 1.31926470 0.187 0.6634467 ## smokeNO -0.07386025 0.941 0.9811578 ## alcabuseNO 0.02501199 0.980 0.9936193 ## bmi -0.17971185 0.857 0.9541543 ## ## Measures for one synthesis and 9 coefficients ## Mean confidence interval overlap: 0.8542318 ## Mean absolute std. coef diff: 0.5714007 ## Lack-of-fit: 5.49732; p-value 0.789 for test that synthesis model is compatible ## with a chi-squared test with 9 degrees of freedom ## ## Confidence interval plot:

While the model needs more work, the same conclusions would be made from both the original and synthetic data set as can be seen from the confidence interavals. Occaisonally there may be contradicting conclusions made about a variable, accepting it in the observed data but not in the synthetic data for example. This scenario could be corrected by using different synthesis methods (see documentation) or altering the visit sequence.
Preserving count data
Released population data are often counts of people in geographical areas by demographic variables (age, sex, etc). Some cells in the table can be very small e.g. <5. For privacy reasons these cells are suppressed to protect peoples identity. With a synthetic data, suppression is not required given it contains no real people, assuming there is enough uncertainty in how the records are synthesised.
The existence of small cell counts opens a few questions,
- If very few records exist in a particular grouping (1-4 records in an area) can they be accurately simulated by synthpop?
- Is the structure of the count data preserved?
To test this 200 areas will be simulated to replicate possible real world scenarios. Area size will be randomly allocated ensuring a good mix of large and small population sizes. Population sizes are randomly drawn from a Poisson distribution with mean ![]() . If large,
. If large, ![]() is drawn from a uniform distribution on the interval [20, 40]. If small,
is drawn from a uniform distribution on the interval [20, 40]. If small, ![]() is set to 1.
is set to 1.
# ---------- AREA
# add area flag to the data frame
area.label <- paste0("area", 1:200)
a <- sample(0:1, 200, replace = TRUE, prob = c(0.5, 0.5))
lambda <- runif(200, 20, 40)*(1-a) + a
prob.dist <- rpois(200, lambda)
area <- sample(area.label, 5000, replace = TRUE, prob = prob.dist)
# attached to original data frame
original.df <- SD2011 %>% dplyr::select(sex, age, socprof, income, marital, depress, sport, nofriend, smoke, nociga, alcabuse, bmi)
original.df$bmi <- ifelse(original.df$bmi > 75, NA, original.df$bmi)
original.df <- cbind(original.df, area) %>% arrange(area)
The sequence of synthesising variables and the choice of predictors is important when there are rare events or low sample areas. If Synthesised very early in the procedure and used as a predictor for following variables, it’s likely the subsequent models will over-fit. Synthetic data sets require a level of uncertainty to reduce the risk of statistical disclosure, so this is not ideal.
Fortunately syn allows for modification of the predictor matrix. To avoid over-fitting, ‘area’ is the last variable to by synthesised and will only use sex and age as predictors. This is reasonable to capture the key population characteristics. Additionally, syn throws an error unless maxfaclevels is changed to the number of areas (the default is 60). This is to prevent poorly synthesised data for this reason and a warning message suggest to check the results, which is good practice.
# synthesise data
# m is set to 0 as a hack to set the synds object and the predictor matrix
synth.obj.b <- syn(original.df, cont.na = cont.na.list, rules = rules.list, rvalues = rules.value.list, maxfaclevels = 200, seed = myseed, m = 0)
##
## Unexpected values (not obeying the rules) found for variable(s): nociga.
## Rules have been applied but make sure they are correct.
# changing the predictor matrix to predict area with only age and sex
new.pred.mat <- synth.obj.b$predictor.matrix
new.pred.mat["area",] <- 0
new.pred.mat["area",c("age", "sex")] <- 1
new.pred.mat
## sex age socprof income marital depress sport nofriend smoke
## sex 0 0 0 0 0 0 0 0 0
## age 1 0 0 0 0 0 0 0 0
## socprof 1 1 0 0 0 0 0 0 0
## income 1 1 1 0 0 0 0 0 0
## marital 1 1 1 1 0 0 0 0 0
## depress 1 1 1 1 1 0 0 0 0
## sport 1 1 1 1 1 1 0 0 0
## nofriend 1 1 1 1 1 1 1 0 0
## smoke 1 1 1 1 1 1 1 1 0
## nociga 1 1 1 1 1 1 1 1 1
## alcabuse 1 1 1 1 1 1 1 1 1
## bmi 1 1 1 1 1 1 1 1 1
## area 1 1 0 0 0 0 0 0 0
## nociga alcabuse bmi area
## sex 0 0 0 0
## age 0 0 0 0
## socprof 0 0 0 0
## income 0 0 0 0
## marital 0 0 0 0
## depress 0 0 0 0
## sport 0 0 0 0
## nofriend 0 0 0 0
## smoke 0 0 0 0
## nociga 0 0 0 0
## alcabuse 1 0 0 0
## bmi 1 1 0 0
## area 0 0 0 0
# synthesising with new predictor
synth.obj.b <- syn(original.df, cont.na = cont.na.list, rules = rules.list, rvalues = rules.value.list, maxfaclevels = 200, seed = myseed, proper = TRUE, predictor.matrix = new.pred.mat)
##
## Unexpected values (not obeying the rules) found for variable(s): nociga.
## Rules have been applied but make sure they are correct.
## Synthesis
## -----------
## sex age socprof income marital depress sport nofriend smoke nociga
## alcabuse bmi area
# compare the synthetic and original data frames
compare(synth.obj.b, original.df, vars = "area", nrow = 1, ncol = 1, cols = c("darkmagenta", "turquoise"), stat = "counts")$plot

The area variable is simulated fairly well on simply age and sex. It captures the large and small areas, however the large areas are relatively more variable. This could use some fine tuning, but will stick with this for now.
tab.syn <- synth.obj.b$syn %>%
dplyr::select(area, sex) %>%
table()
tab.orig <- original.df %>%
dplyr::select(area, sex) %>%
table()
##
## synthetic
## sex
## area MALE FEMALE
## area1 2 0
## area10 15 19
## area101 0 6
## area103 35 22
## area105 3 0
## area106 30 31
## area107 0 3
## area108 28 11
## area110 17 37
## area112 23 24
## area113 0 2
## area114 21 52
## area115 30 28
##
## original
## sex
## area MALE FEMALE
## area1 1 0
## area10 19 27
## area101 1 3
## area103 29 26
## area105 1 0
## area106 22 33
## area107 0 4
## area108 28 18
## area110 18 33
## area112 23 25
## area113 1 2
## area114 28 39
## area115 23 44
d <- data.frame(difference = as.numeric(tab.syn - tab.orig), sex = c(rep("Male", 150), rep("Female", 150)))
ggplot(d, aes(x = difference, fill = sex)) + geom_histogram() + facet_grid(sex ~ .) + scale_fill_manual(values = mycols)
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.

The method does a good job at preserving the structure for the areas. How much variability is acceptable is up to the user and intended purpose. Using more predictors may provide a better fit. The errors are distributed around zero, a good sign no bias has leaked into the data from the synthesis.
tab.syn <- synth.obj.b$syn %>% dplyr::select(marital, sex) %>% table() tab.orig <- original.df %>% dplyr::select(marital, sex) %>% table() ## ## synthetic ## sex ## marital MALE FEMALE ## SINGLE 667 565 ## MARRIED 1352 1644 ## WIDOWED 76 398 ## DIVORCED 81 169 ## LEGALLY SEPARATED 2 0 ## DE FACTO SEPARATED 6 36 ## ## original ## sex ## marital MALE FEMALE ## SINGLE 657 596 ## MARRIED 1382 1597 ## WIDOWED 66 465 ## DIVORCED 62 137 ## LEGALLY SEPARATED 6 1 ## DE FACTO SEPARATED 4 18
At higher levels of aggregation the structure of tables is more maintained.
Build your own methods
‘synthpop’ is built with a similar function to the ‘mice’ package where user defined methods can be specified and passed to the syn function using the form syn.newmethod. To demonstrate this we’ll build our own neural net method.
As a minimum the function takes as input
y– observed variable to be synthesisedx– observed predictorsxp– synthesised predictors
syn.nn <- function (y, x, xp, smoothing, size = 6, ...) {
for (i in which(sapply(x, class) != sapply(xp, class))) xp[,
i] <- eval(parse(text = paste0("as.", class(x[, i]),
"(xp[,i])", sep = "")))
# model and prediction
nn <- nnet(y ~ ., data = as.data.frame(cbind(y, x)), size = size, trace = FALSE)
probs <- predict(nn, newdata = xp)
probs[is.na(probs)] <- 0
for(k in 1:nrow(probs)){
if(sum(probs[k,]) == 0){
probs[k,] <- 1
}
}
new <- apply(probs, 1, function(x) colnames(probs)[sample(1:ncol(probs), 1, prob = x)]) %>% unname()
# if smothing
if (!is.factor(y) & smoothing == "density")
new <- syn.smooth(new, y)
# return
return(list(res = new, fit = nn))
}
Set the method vector to apply the new neural net method for the factors, ctree for the others and pass to syn.
# methods vector
meth.vec <- c("sample", ifelse(sapply(original.df[,-1], is.factor), "nn", "ctree"))
meth.vec[13] <- "ctree"
# synthesise
synth.obj.c <- syn(original.df, method = meth.vec, cont.na = cont.na.list, rules = rules.list, rvalues = rules.value.list, maxfaclevels = 200, seed = myseed, predictor.matrix = new.pred.mat)
##
## Unexpected values (not obeying the rules) found for variable(s): nociga.
## Rules have been applied but make sure they are correct.
## Synthesis
## -----------
## sex age socprof income marital depress sport nofriend smoke nociga
## alcabuse bmi area
# compare the synthetic and original data frames
compare(synth.obj.c, original.df, vars = colnames(original.df)[-13], nrow = 3, ncol = 4, cols = c("darkmagenta", "turquoise"), stat = "counts")$plot
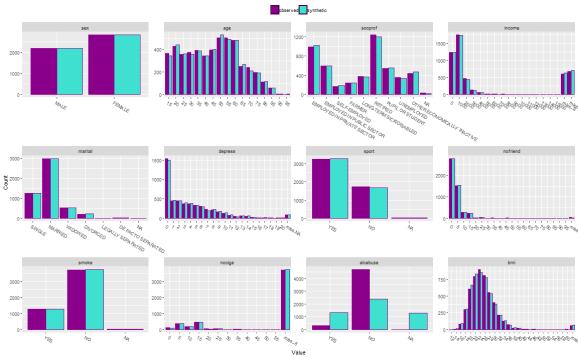
The results are very similar to above with the exception of ‘alcabuse’, but this demonstrates how new methods can be applied.
Takeaways
The ‘synthpop’ package is great for synthesising data for statistical disclosure control or creating training data for model development. Other things to note,
- Synthesising a single table is fast and simple.
- Watch out for over-fitting particularly with factors with many levels. Ensure the visit sequence is reasonable.
- You are not constrained by only the supported methods, you can build your own!
Future posts
Following posts tackle complications that arise when there are multiple tables at different grains that are to be synthesised. Further complications arise when their relationships in the database also need to be preserved. Ideally the data is synthesised and stored alongside the original enabling any report or analysis to be conducted on either the original or synthesised data. This will require some trickery to get synthpop to do the right thing, but is possible.
The post Generating Synthetic Data Sets with ‘synthpop’ in R appeared first on Daniel Oehm | Gradient Descending.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
