Exploring the European Social Survey (ESS) – pipe-friendly workflow with sjmisc, part 2 #rstats #tidyverse
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
This is another post of my series about how my packages integrate into a pipe-friendly workflow. The post focusses on my sjmisc-package, which was just updated on CRAN, and highlights some of the new features. Examples are based on data from the European Social Survey, which are freely available.
Steps of the data analysis process
Let’s recall the last post, where I showed a figure from Hadley Wickham’s R for Data Science book:
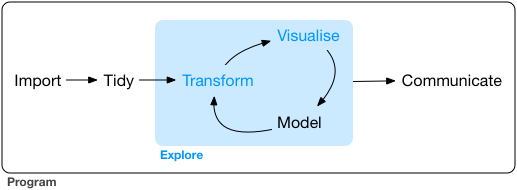
Tidying up, transforming and exploring data is an important part of data analysis, and you can manage many common tasks in this process with the tidyverse- or related packages. The sjmisc-package fits into this workflow, especially when you work with labelled data (like many public use data sets, and the European Social Survey as well), because it offers functions for data transformation and labelled data utility functions. This is what I’m going to show next, based on some „real life“ initial steps when beginning with data exploration.
The European Social Survey
As mentioned above, data from the ESS is freely available. Following examples are based on the SPSS-dataset from the current 7th round. Importing foreign data is easy in R – I think, there’s no other bigger statistical software package, which let’s you import so many data formats.
First, load the required packages and the SPSS-data set. Note that I use the read_spss-function from my sjmisc-package, which wraps the read-function from the haven-package (but it does not return labelled class vectors, for some reasons…). Since the SPSS-data is labelled, the read-function returns a labelled data frame.
library(sjmisc)
library(tidyverse)
ess <- read_spss("ESS7e02.sav")
frq() – printing frequency tables
The first thing that may be of interest might be the response rates in each country. You can plot frequencies for labelled data with the frq()-function. This function requires either a vector or data frame as input and prints the variable label as first line, followed by a frequency-table with values, labels, counts and percentages of the vector.
frq(ess$cntry) # Country val label frq raw.prc valid.prc cum.prc 1 Austria 1795 4.47 4.47 4.47 2 Belgium 1769 4.40 4.40 8.87 3 Switzerland 1532 3.81 3.81 12.68 4 Czech Republic 2148 5.35 5.35 18.03 5 Germany 3045 7.58 7.58 25.60 ... (omitted rows)
find_var() – finding variables in a data frame
Ok, next, let’s look at the distribution of gender across countries. To compute cross tables, you can use the flat_table() function. It requires the data as first argument, followed by any number of variable names. But first, we need to know the name of the gender-variable. It is obviously not „gender“. This is where find_var() comes into play. It searches for variables in a data frame by 1) variable names, 2) variable labels, 3) value labels 4) or any combination of these. By default, it looks for variable name and labels. The function also supports regex-patterns. By default, find_var() returns the column-indices, but you can also print a small „summary“ with the as.varlab-argument.
# find all variables with "gender" in name or label
find_var(ess, "gender", as.varlab = T)
# A tibble: 16 × 3
col.nr var.name var.label
<int> <chr> <chr>
1 150 dscrgnd Discrimination of respondent's group: gender
2 213 icgndra Interviewer code, gender of respondent
3 297 gndr Gender
4 298 gndr2 Gender of second person in household
5 299 gndr3 Gender of third person in household
6 300 gndr4 Gender of fourth person in household
7 301 gndr5 Gender of fifth person in household
... (omitted rows)
Ok, variable in column 297, named „gndr“, is what we are looking for.
flat_table() – printing cross tables
Now we can look at the distribution of gender across countries:
flat_table(ess, cntry, gndr)
# flat_table also works in a pipe-workflow:
ess %>% select(cntry, gndr) %>% flat_table()
gndr Male Female No answer
cntry
Austria 853 942 0
Belgium 896 873 0
Switzerland 766 766 0
Czech Republic 998 1128 0
Germany 1545 1500 0
... (omitted rows)
get_labels() – getting value labels
Next thing we want to know is where to find the educational level of participants. We know that the ESS uses the ISCED-classification. So let’s find „isced“.
find_var(ess, "isced", as.varlab = T) # A tibble: 4 × 3 col.nr var.name var.label <int> <chr> <chr> 1 355 eisced Highest level of education, ES - ISCED 2 447 eiscedp Partner's highest level of education, ES - ISCED 3 496 eiscedf Father's highest level of education, ES - ISCED 4 526 eiscedm Mother's highest level of education, ES - ISCED
ISCED has several categories, but we just want three groups of educational level: low, intermediate and high. So we first need some more information about eisced and its coding. You can get value labels with the get_labels()-function. The simple function call returns the value labels of a variable as character vector, but in this case we also use include.values = "p" to see the related values of the label, and drop.unused = T to drop all labels of those values that don’t occur in the vector.
get_labels(ess$eisced,
include.values = "p",
drop.unused = T)
[1] "[1] ES-ISCED I , less than lower secondary"
[2] "[2] ES-ISCED II, lower secondary"
[3] "[3] ES-ISCED IIIb, lower tier upper secondary"
[4] "[4] ES-ISCED IIIa, upper tier upper secondary"
[5] "[5] ES-ISCED IV, advanced vocational, sub-degree"
[6] "[6] ES-ISCED V1, lower tertiary education, BA level"
[7] "[7] ES-ISCED V2, higher tertiary education, >= MA level"
[8] "[55] Other"
rec() – recoding variables
We just want three categories. The ISCED-classification can be recoded into lower, intermediate and higher educational levels by combining categories 1 and 2, 3 to 5 and 6 and 7. All other values (in the above example, we have a „55“ for „Other“) should be set to missing. We can easily recode and label vectors with the rec()-function. This function does not only recode vectors, it also allows direct labelling of categories inside the recode-syntax (this is optional, you can also use the val.labels-argument). We now recode eisced into a new variable education.
# ok, let's recode
ess$education <- rec(
ess$eisced,
recodes = c("1:2=1 [low]; 3:5=2 [intermediate]; 6:7=3 [high]; else=NA"),
var.label = "Educational level (eisced)"
)
# print frequencies
frq(ess$education)
# Educational level (eisced)
val label frq raw.prc valid.prc cum.prc
1 low 10845 26.99 27.17 27.17
2 intermediate 19978 49.72 50.05 77.21
3 high 9096 22.64 22.79 100.00
4 NA 266 0.66 NA NA
You can see the the vector education has a variable label (Educational level (eisced)), which was set inside the rec()-function, as well as three values with labels („low“, „intermediate“ and „high“). All values 1 to 2 of eisced were recoded into 1 in education, values 3 to 5 into 2 and so on. All remaining values are set to missing (else=NA – for details on the recode-syntax, see ?rec).
Grouped data frames and sjmisc-package
Now, how is education distributed by gender? We can group the ESS-data and print frequencies using the frq()-function for this as well, as this function also accepts grouped data frames. Frequencies for grouped data frames first print the group-details (variable name and category), followed by the frequency table. Thanks to labelled data, the output is easy to understand.
ess %>% select(education, gndr) %>% group_by(gndr) %>% frq() Grouped by: Gender: Male # Educational level (eisced) val label frq raw.prc valid.prc cum.prc 1 low 5023 26.62 26.79 26.79 2 intermediate 9687 51.33 51.66 78.44 3 high 4042 21.42 21.56 100.00 4 NA 119 0.63 NA NA Grouped by: Gender: Female # Educational level (eisced) val label frq raw.prc valid.prc cum.prc 1 low 5813 27.30 27.49 27.49 2 intermediate 10278 48.27 48.61 76.10 3 high 5054 23.74 23.90 100.00 4 NA 147 0.69 NA NA
And what about education and gender across countries? But this time we would like to see the row percentages…
ess %>%
select(gndr, cntry, education) %>%
group_by(gndr) %>%
flat_table(margin = "row")
Grouped by:
Gender: Male
education low intermediate high
cntry
Austria 17.86 69.21 12.93
Belgium 33.93 44.38 21.69
Switzerland 18.19 61.39 20.42
Czech Republic 12.32 74.24 13.43
... (omitted rows)
Grouped by:
Gender: Female
education low intermediate high
cntry
Austria 25.45 61.24 13.31
Belgium 27.68 42.91 29.41
Switzerland 27.58 57.91 14.51
Czech Republic 12.41 75.36 12.23
... (omitted rows)
Nested data frames and models
Next, we want to apply some basic analyses, to see the relation for self-rated health and education in different countries, adjusted for gender and age of respondents. First, we need to find the health-variable. We see that this variable starts with „very good health“ for value 1, and „very bad health“ for value 5. We can reverse this scale, to have better health with higher values, and recode this into a new variable srh (self-rated health):
# find all health-variables find_var(ess, "health", as.varlab = T) # ok, found it. it's named "health". now get coding # of this variable - we see that better health is # coded with lower values get_labels(ess$health) # reverse the scale. there is a "shortcut" pattern # for doing this: "rev" ess$srh <- rec(ess$health, "rev")
So, now we have our dependent variable srh and our independent variables education, gndr and agea.
Let’s get a quick impression about the relations between health and education across countries, by fitting linear models for each country. We can do this using nested data frames (see also this post on bootstrapping with list-variables). The nest()-function from the tidyr package can create subsets of a data frame, based on grouping criteria, and create a new list-variable, where each element itself is a data frame (so it’s nested, because we have data frames inside a data frame).
In the following example, we group the ESS-data by country, and „nest“ the groups. Now we get a data frame with two columns: First, the grouping variable (country) and second, the datasets (subsets) for each country as data frame, store in the list-variable „data“. The data frames in the subsets (in „data“) all contain the selected variables gndr, education, srh and agea.
# convert variable to labelled factor,
# because we then have the labels as
# factor levels in the output
ess$country <- to_label(ess$cntry, drop.levels = T)
ess %>%
select(country, gndr, education, srh, agea) %>%
group_by(country) %>%
nest()
# A tibble: 21 × 2
country data
<fctr> <list>
1 Austria <tibble [1,795 × 4]>
2 Belgium <tibble [1,769 × 4]>
3 Switzerland <tibble [1,532 × 4]>
4 Czech Republic <tibble [2,148 × 4]>
... (omitted rows)
spread_coef() – get back coefficients of nested models
Using the map()-function from the purrr-package, we can iterate this list and apply any function on each data frame in the list-variable „data“. We want to apply the lm()-function to the list-variable, to run linear models for all country datasets. The results of these linear regressions are stored in another list-variable, „models“ (created with the mutate()-function). To quickly access and look at the coefficients, we can use the spread_coef()-function:
ess$country <- to_label(ess$cntry, drop.levels = T)
ess %>%
select(country, gndr, education, srh, agea) %>%
group_by(country) %>%
nest() %>%
mutate(models = purrr::map(data, ~
lm(srh ~ education + gndr + agea,
data = .))) %>%
spread_coef(models)
# A tibble: 21 × 7
country data models `(Intercept)` agea education gndr
<fctr> <list> <list> <dbl> <dbl> <dbl> <dbl>
1 Austria <tibble [1,795 × 4]> <S3: lm> 4.682988 -0.018950774 0.1577497 -0.021005919
2 Belgium <tibble [1,769 × 4]> <S3: lm> 4.325110 -0.012303883 0.1697290 -0.086008553
3 Switzerland <tibble [1,532 × 4]> <S3: lm> 4.435222 -0.011108490 0.1437029 -0.042177216
4 Czech Republic <tibble [2,148 × 4]> <S3: lm> 5.094801 -0.029581163 0.1405284 -0.049518435
5 Germany <tibble [3,045 × 4]> <S3: lm> 4.046202 -0.014770345 0.2021526 -0.051458728
6 Denmark <tibble [1,502 × 4]> <S3: lm> 4.240526 -0.008165113 0.1880305 -0.115486662
7 Estonia <tibble [2,051 × 4]> <S3: lm> 4.098305 -0.023430708 0.2406452 0.003303693
8 Spain <tibble [1,925 × 4]> <S3: lm> 4.613964 -0.018838560 0.1452714 -0.169171898
9 Finland <tibble [2,087 × 4]> <S3: lm> 4.306064 -0.015847129 0.1823691 -0.030827809
10 France <tibble [1,917 × 4]> <S3: lm> 4.143245 -0.013747901 0.2183592 -0.119741127
# ... with 11 more rows
Ok, we see that higher education is positively associated with higher self-rated health, while higher age is negatively correlated to higher self-rated health. What about gender? Forgot the coding of female and male…
get_labels(ess$gndr, include.values = T)
1 2 9
"Male" "Female" "No answer"
Male persons seem to rate their health better than female respondents.
Final words
In this post, I wanted to show a „real life“ approach on exploring unknown datasets. In this example, a labelled dataset was used and shown, how you can work with labelled data in R. The on my sjmisc-package can be useful when performing data analysis, especially in the step of data transformation and exploration. A main focus of this package is to make life with labelled data and data transformation easier, and to integrate into modern workflows (like working with the pipe-operator).
This post demonstrated only a few of the functions the sjmisc-package offers, but I hope you got an impression of its application and functionality.
Tagged: data visualization, R, rstats, sjPlot, Statistik, tidyverse
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
