Fast Bayesian Inference with INLA
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
I am currently a research fellow and 4th year PhD candidate within the INLA group. If you deal with Bayesian models and have never heard about INLA, I sincerely think you should spend a small portion of your time to at least know what it is. If you have heard about it before, you know how nice it is and I would like to ask you to help us spread the word. After all, this can really help some applied researches that work with time-consuming modeling tasks, such as those that involve spatial and spatio-temporal modeling.
INLA is a deterministic method and provides a faster and more accurate alternative to simulation-based MCMC schemes within the class of latent Gaussian models. I am not talking about a slightly improvement over commonly used methods. I am talking about reducing your computational time by orders of magnitude. It is not uncommon to see cases where the computational time got reduced from weeks to hours or from days to minutes.
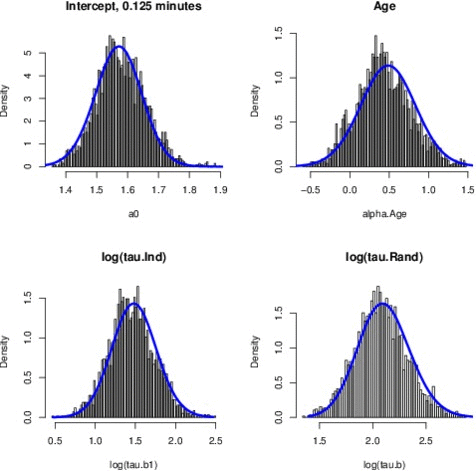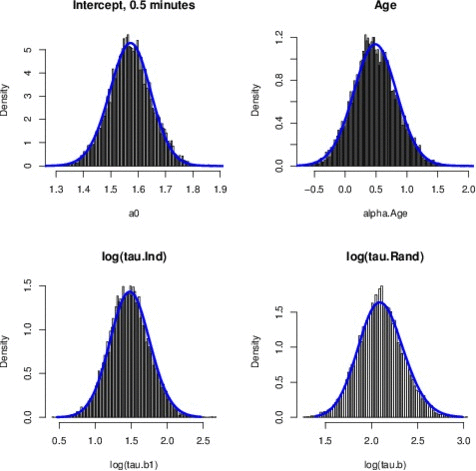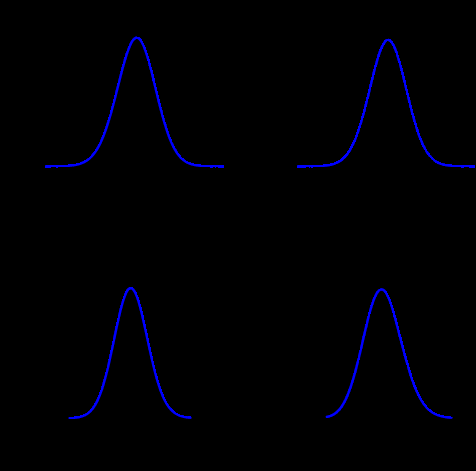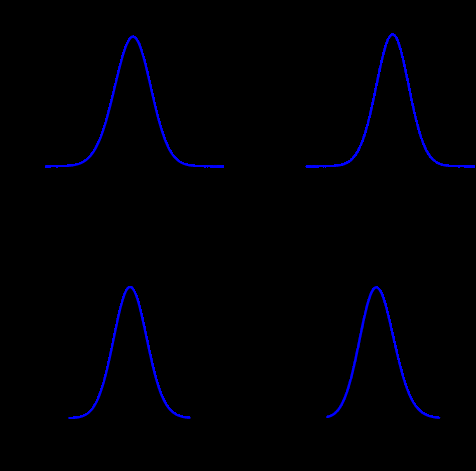 Just to give an example, the Figure above shows the comparison between the posterior marginals approximated by INLA (solid blue line) and MCMC (histograms) for some parameters of the EPIL example. I know the quality of the Figure is not great but those pictures have been computed quite a while ago and I just didn’t have the time to generate those histograms again (remember, I am a 4th year PhD candidate, thesis to finish and all, rs). The approximations returned by INLA are basically instantaneous, while MCMC only match INLA’s accuracy after 16 minutes of computation time (using JAGS). You can find the (very simple) INLA R code for this example in the INLA website.
Just to give an example, the Figure above shows the comparison between the posterior marginals approximated by INLA (solid blue line) and MCMC (histograms) for some parameters of the EPIL example. I know the quality of the Figure is not great but those pictures have been computed quite a while ago and I just didn’t have the time to generate those histograms again (remember, I am a 4th year PhD candidate, thesis to finish and all, rs). The approximations returned by INLA are basically instantaneous, while MCMC only match INLA’s accuracy after 16 minutes of computation time (using JAGS). You can find the (very simple) INLA R code for this example in the INLA website.
As I said before, for large datasets and/or complex models, INLA can be orders of magnitude faster than MCMC. At this point, if you are eager to try INLA I suggest you to download and install the R package INLA and to take a look at the worked out examples in the INLA website. If you stay with me for the weeks to come I plan to write more details and useful information about INLA and its R package. If you are not familiar with the concept of latent Gaussian models, I would like to point out that the following models belong to this broad class:
- Dynamic linear models
- Stochastic volatility
- Generalized linear (mixed) models
- Generalized additive (mixed) models
- Spline smoothing
- Semi-parametric regression
- Space-varying (semi-parametric) regression models
- Disease mapping
- Log-Gaussian Cox-processes
- Model-based geostatistics
- Spatio-temporal models
- Survival analysis
- +++
If you have any doubts regarding INLA there is also a very helpful INLA discussion forum.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
