Faster data exploration with DataExplorer
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.

Data exploration is an important part of the modeling process. It can also take up a fair amount of time. The awesome DataExplorer package in R aims to make this process easier. To get started with DataExplorer, you’ll need to install it like below:
install.packages("DataExplorer")
Let’s use DataExplorer to explore a dataset on diabetes.
# load DataExplorer
library(DataExplorer)
# read in dataset
diabetes_data <- read.csv("https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.csv", header = FALSE)
# fix column names
names(diabetes_data) <- c("number_of_times_pregnant", "plasma_glucose_conc", "diastolic_bp", "triceps_skinfold_thickness", "two_hr_serum_insulin", "bmi", "diabetes_pedigree_function", "age", "label")
# create report
create_report(diabetes_data)
Running the create_report line of code above will generate an HTML report file containing a collection of useful information about the data. This includes:
That’s right – a single line of code can generate all of the above for a given dataset! It’s also possible to get each of these pieces individually. For example, in a single line of code, we can generate histograms for all the numeric variables in the dataset.
plot_histogram(diabetes_data)
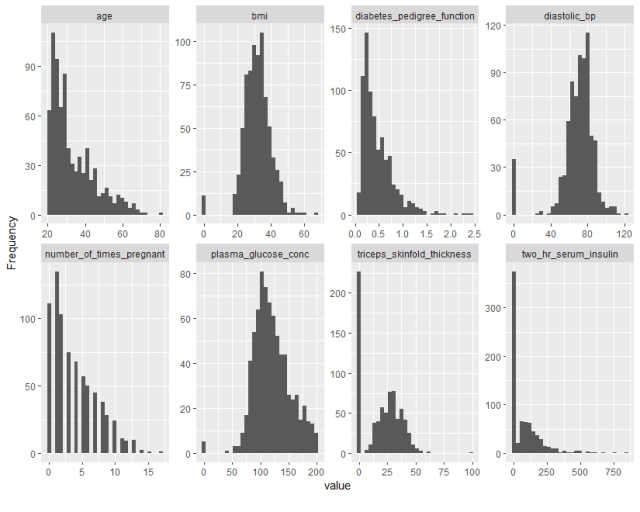
Similarly, we can get bar plots for all categorical variables in the dataset
plot_bar(diabetes_data)
Here’s an example getting the correlation plot:
plot_correlation(diabetes_data)

Configuring the report
It’s also possible to make adjustments to the output generated by create_report. For example, if you don’t want the QQ plots, you could set add_plot_qq = FALSE
config <- configure_report(add_plot_qq = FALSE) create_report(config = config)
One hot encoding
DataExplorer also comes with a function to perform one hot encoding. You can one hot encode all the categorical variables in the dataset by passing the data frame name to the dummify function. In this case, we don’t have any categorical variables to encode, so the function will generate a warning.
dummify(diabetes_data)
Conclusion
That’s all for this post! If you liked this post, consider buying me a coffee. Check out my other R posts by clicking here.
The post Faster data exploration with DataExplorer appeared first on Open Source Automation.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
