Working with Windows CMD system commands in R
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
From time to time, when developing in R, working and wrangling data , preparing for machine learning projects, it comes the time, one would still need to access the operating system commands from/in R.
In this blog post, let’s take a look at some most useful cmd commands when using R. Please note, that the cmd commands apply only to windows environment, for Linux/MacOS, the system commands should be slightly changed, but the wrapper R code should remains the same.

1. R commands to access the CMD
R commands that are used for accessing, running and doing changes through cmd are:
- system
- system2
- shell
- shell.exec
All invoke the OS command that is specified and followed. All of the stated functions are part of the base R, so no additional packages are needed.
Let’s say, we want to start with DIR command and return the results in RStudio.
## 1 Simple dir command with order switch
system("dir /o")
system2("dir /o")
shell("dir /o")
shell.exec("dir /o")
and you will see that shell command will output the results back to console pane in R Studio, where as the others will not give you the immediate desired output. The main difference is that shell function will run the desired command under the shell (following the POSIX standard or here you can find POSIX functions) and the system command will invoke OS command.
In R Base package,a set of dedicated functions is also available to create, open, append and close file and URL connections, as well as opening and loading the compressed files. Just listing the functions:
- object connections: file, url
- compression and encoding: gzfile, bzfile, xzfile, unz, pipe
- connection functions: open,close,flush
- boolean functions: isOpen, isIncomplete
Rule of thumb (with default settings) on when to use system or shell function, shell will work always, system will only work when the system settings (paths) are correctly prepared. Example:
### Checking the usage
Sys.which("copy")
Sys.which("ping")
and the result is that system and system2 functions will work for ping CMD command, but not for copy CMD command, and later in the blogpost, also the rename, move command.

2. Chaining the CMD commands in R
For the simplicity, let’s continue with shell function and try how chaining can be achieved.
Following example will not work, as every time, the shell function is called, the environment is initiated. So running shell sequentially will not work!
setwd("C:\\Users\\Tomaz")
shell("dir")
shell("cd ..")
shell("dir")
Chaining the cmd commands together in one command will work:
shell("dir && cd .. && dir")
Note that this will not alter your working directory, so every time the shell is completed, you will be returned back to initiated environment. By checking:
getwd()
you can see, that the working directory has not changed. Checking before and after the R environment variables, these will also remained intact:
Sys.getenv(c("R_HOME","HOME"))
You can also define the variable, use the echo command, use the build in system variables like %TIME% and many more. More is available at Github.
3. Most typical CMD commands executed from R
Here is the list of some CMD commands we will encapsulate in R:
- fc / compare
- rename
- move / copy
- ping (pingpath)
- systeminfo
- tasklist (taskkill)
- ipconfig / netstat
3.1. Comparing two files in CMD
Having capability to compare two files in CMD is just another of many ways to check if your dataset is the same.
Let’s create two *.csv files with iris dataset and run the file compare in cmd:
setwd("C:\\Users\\Tomaz")
#create two files
write.csv(iris, file="iris_file1.csv")
write.csv(iris, file="iris_file2.csv")
#compare both files
shell("FC /a C:\\Users\\Tomaz\\iris_file1.csv
C:\\Users\\Tomaz\\iris_file2.csv")
The result information is: FC: no differences encountered. Same as if the command would be run from command line:

Same result could be achieved with R function that comes in base build: all.equal.
file1 <- "C:\\Users\\Tomaz\\iris_file1.csv" file2 <- "C:\\Users\\Tomaz\\iris_file2.csv" # or in R using all.equal all.equal(readLines(file1), readLines(file2))
Since we are in R, we would like to have results stored in data.frame. To do so, let’s use the previously mentioned System2 function. This is a newer version of System function that uses arguments and commands separately and some additional settings.
# run the same command using newer function "system2" and set the arguments
cmd_command <- "FC"
#cmd_args <- "/a C:\\Users\\Tomaz\\iris_file1.csv
C:\\Users\\Tomaz\\iris_file2.csv"
cmd_args <- c('/a', file1, file2)
rr <- system2(command=cmd_command,
args= cmd_args,
stdout=TRUE,
stderr=TRUE,
wait = TRUE)
#suppose we want to store the results in data.frame
#empty dataframe
df_rr <- data.frame(file1 = character(),
file2 = character(),
fcompare = character(),
stringsAsFactors=FALSE)
#temporary results
temp <- data.frame(file1=file1, file2=file2, fcompare=rr[2])
#bind all into dataframe
df_rr <- rbind(df_rr, setNames(temp, names(df_rr)))
with results stored in data.frame when comparing multiple files.

3.2. Renaming a file in CMD
Keeping the file with iris dataset: C:\\Users\\Tomaz\\iris_file1.csv, let’s do the renaming.
# renaming the file using shell command
shell("cd c:\\users\\tomaz && ren iris_file1.csv iris_file1_copy.csv")
and with the parametrized shell function (remember the copy will not work due to Sys.which(“copy”) is returned empty):
file_old_name <- "c:\\users\\Tomaz\\iris_file2.csv"
file_new_name <- "iris_file2_new.csv"
cmd_command <- paste("RENAME", file_old_name, file_new_name)
# System2 Does not work
# system2(command=cmd_command)
shell(cmd_command)
In both ways, the files are renamed. Fortunately or not, the output in this case is none, when there is no syntax error and file exists.
3.3. Copy a file in CMD
Keeping the file with iris dataset: C:\\Users\\Tomaz\\iris_file1.csv, let’s do also the copying.
#Copying file
shell("copy c:\\Users\\Tomaz\\iris_file1.csv iris_file1_copy.csv")
and with the parametrized shell function:
orig_file <- "c:\\Users\\Tomaz\\iris_file1.csv" copy_file <- "iris_file1_copy1.csv" command <- "copy" cmd_command <- paste(command, orig_file, copy_file) shell(cmd_command)
Now check your files.
3.4. Ping
Since ping CMD is available with the sys.which() function, we can use it directly as:
URL <- 'tomaztsql.wordpress.com'
cmd_command <- "Ping"
cmd_args <- c('-n 1', URL)
system2(command=cmd_command,
args=cmd_args,
stdout=TRUE,
stderr=TRUE,
wait = TRUE)
And since we want to check many URLs, let’s create an empty data.frame to store the result and a function to walk through the list.
URLs <- c("google.com", "tomaztql.wordpress.com", "youtube.com")
#empty dataframe
df_rr <- data.frame(URL = character(),
reply = character(),
package = character(),
stringsAsFactors=FALSE)
ping_fn <- function(url) {
system2("ping",c(url,' -n 1'),
stdout=TRUE,
stderr=TRUE)
}
for (i in 1:length(URLs)){
site <- print(URLs[i])
rr <- ping_fn(site)
temp <- data.frame(URL=rr[2], reply=rr[3], package=rr[6])
df_rr <- rbind(df_rr, setNames(temp, names(df_rr)))
}
head(df_rr)
and the result is binded in a single data.frame:

3.5. Systeminfo
Assume, you need the information about the system not only about the R environment. Some of the R functions are available here:
# R R.Version() Sys.info() .Platform
And all are very useful. But if you need Systeminfo CMD information into your R IDE environment, save it in data.frame, using system2() function is a way to go.
# Using system2 cmd_command <- "systeminfo" rr <- system2(command=cmd_command, stdout=TRUE, stderr=TRUE, wait = TRUE)
and getting the result:
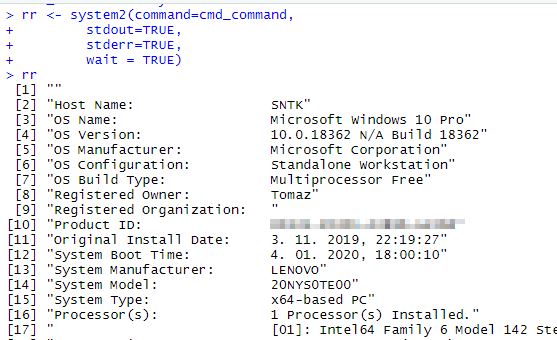
So grabbing the e.g.: “System Manufacturer” just walk to the 13th position in the vector:
rr[13]
3.6. Tasklist
Getting the list of all the running task
cmd_command <- "tasklist"
rr <- system2(command=cmd_command,
stdout=TRUE,
stderr=TRUE,
wait = TRUE)
# getting the results into something more readable format
dd <- tibble::enframe(rr)
stack(dd)[1]
and can get all the additional information from the data.frame.
In addition, you can also kill a specific task
# kill a specific task
shell("taskkill /F /PID 6816")
with the returned information: “SUCCESS: Sent termination signal to the process with PID 6816.“
3.7 Ipconfig / netstat
Getting the list IpConfig information or all the netstat information, the task is the same:
cmd_command <- "netstat"
rr <- system2(command=cmd_command,
stdout=TRUE,
stderr=TRUE,
wait = TRUE)
and storing the results with tibble:
dd <- tibble::enframe(rr)
and the results are same as in CMD:

4. Conclusion
Using CMD in R IDE will for sure not be your daily driver when crunching numbers and wrangling data. But when you will need to operate closely with your files, datasets, system or tasks, system2 and shell functions are absolute winners to do the job.
As always, complete set of the code is available on Github.
Happy R-coding! 
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
