Finding duplicates in data frame across columns and replacing them with unique values using R
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
Suppose you have a dataset with many variables, and you want to check:
- if there are any duplicated for each of the observation
- replace duplicates with random value from pool of existing values.
In this manner, let’s create a sample dataset:
df <- structure(list(
v1 = c(10,20,30,40,50,60,70,80)
,v2 = c(5,7,6,8,6,8,9,4)
,v3 = c(2,4,6,6,7,8,8,4)
,v4 = c(8,7,3,1,8,7,8,4)
,v5 = c(2,4,6,7,8,9,9,3))
,.Names = c("ID","a", "b","d", "e")
,.typeOf = c("numeric", "numeric", "numeric","numeric","numeric")
,row.names = c(NA, -8L)
,class = "data.frame"
,comment = "Sample dataframe for duplication example")
which has the following interesting characteristics:

Upon closer inspection, one will see that there are many duplicated values across different variables (variable ID, variable a, variable b, variable d and variable e). So let’s focus on:
- row 2 has two times duplicated values (2x value 4 and 2x value 7)
- row 3 has three times duplicated values (3x value 6)
Our pool of possible replacement values are:
possible_new_values <- c(1,2,3,4,5,6,7,8,9
Creating loop for slicing the data, loop through the duplicated positions, at the end looks like:
for (row in 1:nrow(df)) {
vec = df %>% slice(row) %>% unlist() %>% unname()
#check for duplicates
if(length(unique(vec)) != length(df)) {
positions <- which(duplicated(vec) %in% c("TRUE"))
#iterate through positions
for(i in 1:length(positions)) {
possible_new_values <- c(1,2,3,4,5,6,7,8,9)
df[row,positions[i]] <- sample(possible_new_values
[ ! possible_new_values %in% unique(vec)],1)
}
}
}
revealing the final replacement of values in

So the end result, when putting old data.frame and the new data.frame (with replaced values) side by side, it looks like:
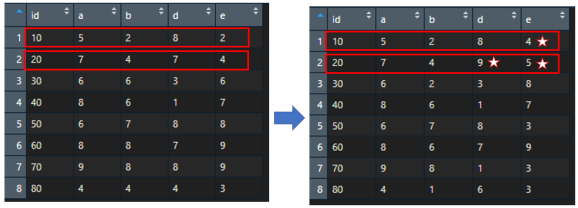
Showing how replacement works per each row across given columns | rows.
Niffy, yet useful data de-duplication or data replacements, when you need one.
As always, code is available at Github.
Happy coding with R 
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
