Beads Summary Plot of Ranges for R
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
The beadsplot function is designed for a data frame with a factor column and many observation columns. This function summarize the data visually. The builtin iris data is suitable to start with this function quickly, because it has a factor on 5th column, and other 4 columns are numeric observations.
Let’s make a summary table of iris data first, without using the beadsplot.
> str(iris)
'data.frame': 150 obs. of 5 variables:
$ Sepal.Length: num 5.1 4.9 4.7 4.6 5 5.4 4.6 5 4.4 4.9 ...
$ Sepal.Width : num 3.5 3 3.2 3.1 3.6 3.9 3.4 3.4 2.9 3.1 ...
$ Petal.Length: num 1.4 1.4 1.3 1.5 1.4 1.7 1.4 1.5 1.4 1.5 ...
$ Petal.Width : num 0.2 0.2 0.2 0.2 0.2 0.4 0.3 0.2 0.2 0.1 ...
$ Species : Factor w/ 3 levels "setosa","versicolor",..: 1 1 1 1 1 1 1 1 1 1 ...
> lapply(list(max=max, mean=mean, min=min), function(FUN)
apply(iris[1:4], 2, function(x)
tapply(x, iris[,5], FUN)))
$max
Sepal.Length Sepal.Width Petal.Length Petal.Width
setosa 5.8 4.4 1.9 0.6
versicolor 7.0 3.4 5.1 1.8
virginica 7.9 3.8 6.9 2.5
$mean
Sepal.Length Sepal.Width Petal.Length Petal.Width
setosa 5.006 3.428 1.462 0.246
versicolor 5.936 2.770 4.260 1.326
virginica 6.588 2.974 5.552 2.026
$min
Sepal.Length Sepal.Width Petal.Length Petal.Width
setosa 4.3 2.3 1.0 0.1
versicolor 4.9 2.0 3.0 1.0
virginica 4.9 2.2 4.5 1.4
Someone may say this is enough, but I want more visuals.
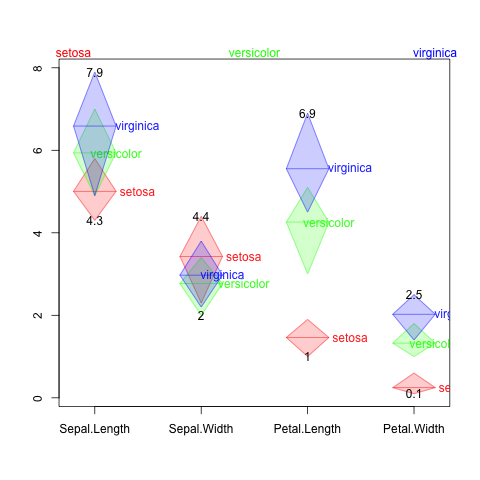
beadsplot(Species~., iris, scale.mean=NULL, scale.range=NULL)
Observations may have different ranges and units between columns. Eg. soil chemical component data frame may have pH, EC, Lime, Phosphoric acid, CEC and so on. These columns have quite different units. So I need a way to separate y-axis scales by column. Let’s do the relative scaling by the whole range of each column.
beadsplot(Species~., iris, scale.mean=NULL)
In Fig. 2, the y-axis has no units, and every column is scaled between -1 and 1.
Another relative scaling is using the whole mean with the range. The mean of each column is also a useful information.
beadsplot(Species~., iris)
Fig. 3 is very similar to Fig.2. But the y-axis is slightly shifted at each column. Now the mean of each column is located at y=0. It is showing asymmetry of ranges.
The beadsplot can be used to show summary table like the first one.
beadsplot(Species~., iris, plot=FALSE) $scaled , , summaries = S factors series setosa versicolor virginica Sepal.Length -0.8574074 -0.5240741 -0.5240741 Sepal.Width -0.6311111 -0.8811111 -0.7144444 Petal.Length -0.9349153 -0.2569492 0.2515254 Petal.Width -0.9161111 -0.1661111 0.1672222 , , summaries = E factors series setosa versicolor virginica Sepal.Length -0.4651852 0.05148148 0.41370370 Sepal.Width 0.3088889 -0.23944444 -0.06944444 Petal.Length -0.7783051 0.17016949 0.60813559 Petal.Width -0.7944444 0.10555556 0.68888889 , , summaries = N factors series setosa versicolor virginica Sepal.Length -0.02407407 0.6425926 1.1425926 Sepal.Width 1.11888889 0.2855556 0.6188889 Petal.Length -0.62983051 0.4549153 1.0650847 Petal.Width -0.49944444 0.5005556 1.0838889 attr(,"summary.labels") S ".Primitive(\"min\")" E "function (x, ...) UseMethod(\"mean\")" N ".Primitive(\"max\")" $raw , , summaries = S factors series setosa versicolor virginica Sepal.Length 4.3 4.9 4.9 Sepal.Width 2.3 2.0 2.2 Petal.Length 1.0 3.0 4.5 Petal.Width 0.1 1.0 1.4 , , summaries = E factors series setosa versicolor virginica Sepal.Length 5.006 5.936 6.588 Sepal.Width 3.428 2.770 2.974 Petal.Length 1.462 4.260 5.552 Petal.Width 0.246 1.326 2.026 , , summaries = N factors series setosa versicolor virginica Sepal.Length 5.8 7.0 7.9 Sepal.Width 4.4 3.4 3.8 Petal.Length 1.9 5.1 6.9 Petal.Width 0.6 1.8 2.5 attr(,"summary.labels") S ".Primitive(\"min\")" E "function (x, ...) UseMethod(\"mean\")" N ".Primitive(\"max\")" $scale $scale$scale.range [1] 1 $scale$scale.mean [1] 0 $scale$scale.log [1] FALSE $scale$scale.data.center NULL $scale$scale.data.border NULL $scale$scale.grid.center [1] "grey" $scale$scale.grid.border [1] "grey" $scale$cex.axis [1] 1
Download is available at http://code.google.com/p/cowares-excel-hello/downloads/list?q=label:diaplt_r .
Expected to be available at CRAN soon.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.