Data Profiling in R
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
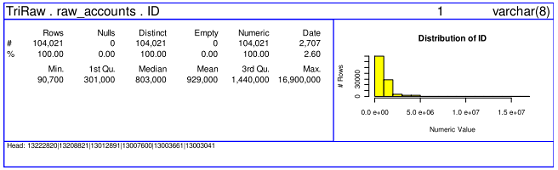
In 2006 UserR conference Jim Porzak gave a presentation on data profiling with R. He showed how to draw summary panels of the data using a combination of grid and base graphics.
Unfortunately the code has not (yet) been released as a package, so when I recently needed to quickly review several datasets at the beginning of an analysis project I started to look for alternatives. A quick search revealed two options that offer similar functionality: r2lUniv package and describe() function in Hmisc package.
r2lUniv
r2lUniv package performs quick analysis either on a single variable or on a dataframe by computing several statistics (frequency, centrality, dispersion, graph) for each variable and outputs the results in a LaTeX format. The output varies depending on the variable type.
> library(r2lUniv) |
One can specify the text to be inserted in front of each section.
> textBefore <- paste("\\subsection{", names(mtcars),
+ "}", sep = "")
> rtlu(mtcars, "fileOut.tex", textBefore = textBefore)
|
The function rtluMainFile generates a LaTeX main document design and allows to further customise the report.
> text <- "\\input{fileOut.tex}"
> rtluMainFile("r2lUniv_report.tex", text = text)
|
The resulting tex-file can then be converted into pdf.
> library(tools)
> texi2dvi("r2lUniv_report.tex", pdf = TRUE, clean = TRUE)
|
A sample output for the mpg-variable:

The final pdf-output can be seen here: r2lUniv_report.pdf.
Hmisc
The describe function in Hmisc package determines whether the variable is character, factor, category, binary, discrete numeric, and continuous numeric, and prints a concise statistical summary according to each. The latex report also includes a spike histogram displaying the frequency counts.
> library(Hmisc) |
> db <- describe(mtcars, size = "normalsize") |
The easiest and fastest way is to print the results to the console.
> db$mpg
mpg
n missing unique Mean .05 .10 .25 .50
32 0 25 20.09 12.00 14.34 15.43 19.20
.75 .90 .95
22.80 30.09 31.30
lowest : 10.4 13.3 14.3 14.7 15.0
highest: 26.0 27.3 30.4 32.4 33.9
|
Alternatively, one can convert the describe object into a LaTeX file.
> x <- latex(db, file = "describe.tex") |
cat is used to generate the tex-report.
> text2 <- "\\documentclass{article}\n\\usepackage{relsize,setspace}\n\\begin{document}\n\\input{describe.tex} \n\\end{document}"
> cat(text2, file = "Hmisc_describe_report.tex")
|
> library(tools)
> texi2dvi("Hmisc_describe_report.tex", pdf = TRUE)
|
A sample output for the mpg-variable:

The final pdf-report can be seen here: Hmisc_describe_report.pdf.
Conclusion
Both of the functions provide similar snapshots of the data, however I prefer the describe function for its more concise output, and also for the option to print the analysis to the console. Whilst I like the summary plots generated by r2lUniv I find them hard to read in the pdf-report because of the small font-size of the labels.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
