from OTU table to HEATMAP!
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
In this tutorial you will learn:
- what is a heatmap
- how to create a clean, highly customizable heatmap using heatmap.2 in the gplots package in R
- how to remove samples with poor output (not very many sequences)
- how to rearrange your samples by a metadata category
- how to make a color coded bar above the heatmap for the metadata category
You will not learn:
- how to create an OTU table
- how to choose a good color palette for your color coded bar
- why heatmaps are called heatmaps
What you will need:
- metadata table
- OTU table
- and in case it wasn’t obvious, R
Introduction
So figuring out a code from OTU table to heatmap has been my dream since we saw a cool looking heatmap in one of Dr. Lamendella’s presentations on the human gut microbiome (from the most awesome gut microbiome paper ever of 2012). It is a neat way to display a matrix of information in a color coded grid and is not in any way related to Fahrenheit or Celsius. One of the most common applications of heatmaps are for displaying results of gene expression levels in DNA microarrays.

Photo from Wikipedia: Heatmap of a DNA microarray-Unfortunately for people with red-green color-blindness, these two colors are the most commonly used for creation of heatmaps.
In our case, the heatmap will function particularly well in displaying information about OTU abundances at various taxonomical levels. Heatmaps would also serve really well when people in the lab start theoretically getting metaproteomic data. To see more examples of heatmaps used in seminal research, look at the supplementary figures of the coolest paper of 2012. This is a particularly long tutorial that has codes adapted from several sources. I will try my best to explain each step but some of the steps in certain functions are still a bit foggy and others would require unnecessary and long explanations. My original bare basic tutorial on heatmaps can be found here and it was mainly based upon this tutorial done by Nathan Yu. If you are keen on just making a quick heatmap, I suggest looking at those two links first.
Prepping the Files
You know how it goes, you need to make things in the right format before you can make the magic happen.
- Metadata File-Your sample names should listed down the first column and variables describing your samples are in the top row. You can have as many columns/variables as you like.
- OTU Table-Contrary to the metadata file, your sample names should be on the top row whereas the OTU IDs should be down the first column. When you scroll all the way to the right on your file there should be a column with the taxonomical information in each row such as
- “k__Bacteria; p__Firmicutes; c__Clostridia; o__Clostridiales; f__Lachnospiraceae; g__Coprococcus; s__Coprococcus eutactus” or
- “Root;Bacteria;Firmicutes;””Clostridia””;Clostridiales;””Lachnospiraceae””;””Lachnospiraceae Incertae Sedis”
-
Unlike the columns which have a title (the sample name), this one is probably blank. Type in “taxonomy”. All lowercase and no extra frills.
-
Make sure the ID numbers/names for samples MATCH between your metadata file and OTU file.
- In reality, it will probably make things easier for you if your samples are something like “Site2Samp3Vers1″ rather than just “231″. This is one of those critical steps that absolutely must be done or else R won’t be able to match up the OTU data of each sample with the metadata of each respective sample. When creating your sample names, the concatenation will probably help you out as explained here.
-
Make sure both are saved as .csv (Comma-Separated Values) rather than .xls or .xlsx rather than Excel.
- This compacts the file and makes the importation process into R easier.
Hopefully this didn’t take you too long! Now the fun begins.
Importing the files into R
OPTIONAL: I tend to have stuff saved accidentally in my R sessions that I no longer need, so my first step of every large script is removing anything that is currently in the work session.
ls() rm(list=ls()) ls()
OPTIONAL: I also usually change my working directory at the beginning. This makes writing pathways (or the directions for R to find files) much easier and leaves less of a chance for things to get messed up. Write in the pathway for the folder where your metadata and OTU tables are located. The one written below follows the pathway conventions of Windows PC (for Mac view http://youtu.be/8xT3hmJQskU).
getwd()
setwd("c:/your_folder/project_1")
getwd()
OPTIONAL: The second getwd() call should repeat whatever working directory you just specified. To double check that your files are in the pathway you just specified:
dir()
Now we actually import the files into R. Change the names to whatever you named your file.
dat1 <- read.csv("OTU.csv", header=TRUE,row.names=1, sep=",")
meta <- read.csv("metadata.csv", header=TRUE,row.names=1, sep=",")
Also take the time to make a folder called “output” in your working directory folder. We will be writing lots of scripts that spit out tables here and there, so it is nice to separate them out. What are you looking for? Just do it manually (you know, right click, create new folder, etc.); it’s easier.
OTU Table “Manipulation” or Cleaning
As you already know, data usually don’t arrive in the ready to analyze format. From missing values to messed up samples, now we take the time to clean our data (since the phrase “data manipulation” is not PC at all). In this process, we will split the OTU table into two segments/objects. The first object (taxa.names) will contain just the taxonomical information and the second object is a matrix(dat2) that will contain the rest of the table.
taxa.names <- dat$taxonomy dat2 <- as.matrix(dat[,-dim(dat)[2]])
Optional: Dropping Samples with Low Observations
This section is adapted from Dr. Schaefer’s tutorial “Analyses of Diversity, Diversity and Similar Indices” class notes. The whole class syllabus is rather fascinating and proves a lot of R equivalents of processes (Unifrac analysis, nMDS, Indicator Species analysis, etc.) that we would normally do in QIIME or PC-ORD.
Although the subtitle says optional, it’s probably a good idea to to drop samples with low observations. There is a very low probability your sample only has 10 or even just 100 microbes in it.
First, create an object that contains number of observances for each sample (sums each column).
s_abundances<-apply(dat2,2,sum)
Next, split the OTU table into two matrices: one with low abundances and one with high abundances. Currently the threshold is set at 1000 but you can change the number in both lines.
bads<-dat2[,s_abundances<1000] goods<-dat2[,s_abundances>1000]
To see how many poor samples will be removed from the OTU table:
ncol(dat2)-ncol(goods) ncol(bads)
Now replace the old OTU table with the new one that just contains good markers.
dat2<-goods
Run these if you would like output of the good and bad markers.
write.table(badm, file="output/bads.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE) write.table(goodm, file="output/goods.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE)
.
The next parts on changing to relative abundance and extraction by different taxonomic levels are based on a powerpoint in the lab module “Introduction to Metagenomic Data Analysis” by Alexander V. Alekseyenko and Susan P. Holmes (as part of the Summer Institute in Statistics and Modeling of Infectious Diseases).
Recommended: Change to Relative Abundance
Right now, observances of each OTU are probably being reported as absolute abundances. Each OTU observance is the original number generated by OTU picking and the like in QIIME. Sometimes there are specific reasons why you might want to leave the abundances as absolute abundances but the majority of the time, you will probably need to make samples comparable to each other. This requires standardization: the absolute number of observances for an OTU becomes a fraction of the total observances in that sample.
dat2 <- scale(dat2, center=F, scale=colSums(dat2))
To begin, you will need transpose the OTU table so that the OTU ID numbers become the top row rather than the first column.
dat2 <-t(dat2)
Next, take a closer look at the way taxonomical designations are written in your OTU table. Are there six or seven levels? If it looks like
- “Root;Bacteria;Firmicutes;”"Clostridia”";Clostridiales;”"Lachnospiraceae”";”"Lachnospiraceae Incertae Sedis”
- “k__Bacteria; p__Firmicutes; c__Clostridia; o__Clostridiales; f__Lachnospiraceae; g__Coprococcus; s__Coprococcus eutactus”
there are seven levels designated by the Root/Kingdom, Phylum, Class, Order, Family, Genus and Specie. You will need to start numbering the levels at level 2 rather than 1 in the extraction functions. However, if the beginning of your taxonomical designations are phyla then start numbering at level 1; or if phyla are not listed until the third spot in the taxonomical designations, start numbering the levels at 3.
d.phylum = otu2taxonomy(dat4,level=2,taxa=taxa.names) d.class = otu2taxonomy(dat4,level=3,taxa=taxa.names) d.order = otu2taxonomy(dat4,level=4,taxa=taxa.names) d.family = otu2taxonomy(dat4,level=5,taxa=taxa.names) d.genus = otu2taxonomy(dat4,level=6,taxa=taxa.names) d.species = otu2taxonomy(dat4,level=7,taxa=taxa.names)
To look at the output, transpose and export.
phylum2 <-t(d.phylum) class2 <-t(d.class) order2 <-t(d.order) family2 <-t(d.family) genus2 <-t(d.genus) species2 <-t(d.species) write.table(phylum2, file="output/phyla.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE) write.table(class2, file="output/classes.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE) write.table(order2, file="output/orders.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE) write.table(family2, file="output/families.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE) write.table(genus2, file="output/genera.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE) write.table(species2, file="output/species.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE)
These steps allow you to analyze your OTU data at various taxonomical levels within and outside of R.
Merging metadata and OTU tables
Next we will merge the metadata and OTU table together. Originally, I tried to figure out a way to avoid this step but R was not being so cooperative with reordering. We are merging the two tables so that when we rearrange the samples according to a metadata variable, the OTU table rearranges as well.
First choose which taxonomical level you want to look at. For simplicity’s sake, let’s look at phyla first. This first script merges the two tables by matching up the Sample ID/Names. The “all.x=FALSE” portion makes sure that metadata information is not included for samples that were dropped earlier from the data processing (if you skipped that step, this should not affect the output).
mergephylum<-merge(meta, d.phylum, by="row.names",all.x=FALSE)
There are multiple ways to double check the merge worked. The easiest is just by looking at the dimensions of each object involved.
dim(meta) dim(d.phylum) dim(mergephylum)ncol
The first number reflects the number of rows while the second refers to the amount of columns. The number of rows of the final merge should match the number of rows in the phyla file (or whichever level you are looking at). The number of columns of the final merge should be the sum of the columns in d.phylum and meta +1. Again, here is the code for output.
write.table(mergephylum, file="output/mergephylum.csv", col.names=NA,row.names=TRUE, sep=",", quote=FALSE)
Reordering by metadata variable
If you would like your heatmap in order by sample or order does not matter, skip to the next step (making the actual matrix for the heatmap).
You can reorder your samples based on a variable in your metadata. This can be anything from diet type to sample site. The variable that will direct the way your table will be reordered is not limited to just being discrete (as in group A, group B, etc.); the variable can also be continuous (blood pressure, height, pH).
For this rearrangement code, you will need the exact name of the column of the variable. Since names might have changed since importation into R, it’s best to double check. This lists the names of all the columns of your metadata.
colnames(meta)
Now use the name of the variable in question and replace “VARIABLE” in the following code.
reordermerge <- mergephylum[order(mergephylum$VARIABLE, mergephylum$Row.names),]
This will cause the table to be rearranged by increasing value of the VARIABLE (or alphabetical order) and then by the Row.names or Sample ID/Names.
Making the actual matrix for the heatmap
This next set of script splits OTU table and metadata again so that the heatmap will only display OTU data rather than metadata as well. It involves removing columns, renaming the rows and changing the format of the object to a matrix.
(OTUcol1<-ncol(meta)+2) (OTUcol2<-ncol(reordermerge)) justOTU<-reordermerge[,OTUcol1:OTUcol2] justOTU[1:5,1:5] rownames(justOTU[1:10,]) rownames(justOTU)<-reordermerge$Row.names rownames(justOTU[1:10,]) justOTU2<-as.matrix(t(justOTU)) justOTU2[1:5,1:5]
Heatmap!!!
Hurray! After all those steps we are finally to the steps of making the actual heatmap.
First install the package that contains the codes to make the heatmap. (Refresh your memory on packages here.)
install.packages("gplots")
library(gplots)
Now print out your heatmap!
heatmap.2(justOTU2,Rowv=TRUE, Colv=FALSE, scale="column", trace="none", col=redgreen, xlab="sample", ylab="phylum", margins=c(10,15))
If you are happy with this image, skip to “Export the Heatmap.“
This tutorial will be covering some of the arguments within this function for customization. In case you want to know about all the possibilities, read this documentation on heatmap.2.
Looking at some of the arguments already present:
- justOTU2-This is where you put the object name of the heatmap you would like to visualize.
- Colv and Rowv-Hold your horses and read about these in the next subsection.
- scale-For the scale argument you can input “none”, “column” or “row”. R normalizes the rows or columns based on the Z-score or standard score (not to be confused with Z-factor) which indicates how many standard deviations away the value is from the mean. In other words, the scale indicates which values are outliers and possibly significant.
- trace-You should probably set this to “none” because if you put “column” “row” or “both”, you get some funky lines separating the boxes in the heatmap. Unless your data set is pretty small, then you would benefit from adding the following code with a different color.
-
tracecol="cyan"
-
- col-This controls the color scheme. Read about changing the color scheme in the second following subsection.
- xlab and ylab-This is where you put labels for your X and Y axis.
- margins-Changing the numbers in this argument changes the amount of white space surrounding your heatmap. I highly recommend fiddling with these numbers to make sure the visualization space fits the heatmap of your data.
Optional: Dendrograms
A dendrogram is a tree placed on right and/or top sides of the heatmap. Instead of clustering phylogenetically similar samples or phyla, these trees cluster columns or rows that have similar values. This clustering causes the rows and columns to be reordered when you input “TRUE” for Rowv or Colv. If you input Colv=TRUE, then you will lose any customized ordering you did earlier when you reordered the metadata.
Optional: Change the color scale
Although many default heatmaps use the red-green color scheme, I would suggest choosing a different palette to A) better match the color scheme of your paper and more seriously, B) allow color blind people the chance to understand your graphic.
I will not go much into the specifics of choosing good color palettes, but the most important aspect of designing graphics is that people can actually read and understand them. The color scheme must assist in this goal. So yellow text on a white background is a huge no-no (regardless if the audience is color blind or not) and remember, there are people who can be blue-green color blind. In case you want to learn more about colors in R in general, I stumbled across this presentation that has excellent information but is displayed in the most horrendous manner graphically.
There are already a couple color schemes present in R. You can find them by entering
?cm.colors
The easiest way to change the color scheme to a customized scheme is by using the package “rcolorbrewer.”
install.packages("RColorBrewer")
library(RColorBrewer)
The information in your heatmap would probably benefit best from a monochromatic color scheme (white, light blue, dark blue, etc.) or a diverging color scheme (dark red, pink, white, light blue, dark blue, etc.) Creating both utilize a similar function from the RColorBrewer: colorRampPalette. (For more information about RColorBrewer, look at an explanation here and and another one by Dr. Stewart MacArthur).
For a monochromatic scale, change the color names within the quotation markers:
scaleblue<-colorRampPalette(colors=c("white","blue"))(100)
For a diverging scale, input three colors.
scaleyellowblue<-colorRampPalette(colors=c("yellow","white","blue"))(100)
The number at the end corresponds to the number of colors generated in the scale. Thus if you inputted
scaleyellowblue<-colorRampPalette(colors=c("yellow","white","blue"))(3)
You would only get the 3 original colors in the scale.
To apply the color scale, input the name of your custom scale in the argument for col.
heatmap.2(justOTU2,Rowv=TRUE, Colv=FALSE, scale="column", trace="none", col=COLORSCALE, xlab="sample", ylab="phylum", margins=c(10,15))
For example, the diverging yellow-white-blue color scheme:
heatmap.2(justOTU2,Rowv=TRUE, Colv=FALSE, scale="column", trace="none", col=scaleyellowblue, xlab="sample", ylab="phylum", margins=c(10,15))
If you want to get fancy, try choosing colors from:
For example, the Science paper I mentioned earlier had a really cool protein heatmap with a customized color scheme. To replicate that scheme:
colors in R
- http://simplystatistics.org/2011/10/17/colors-in-r/
- http://www.compbiome.com/2010/12/r-using-rcolorbrewer-to-colour-your.html
Optional: Colored Side Bar
In the several papers, you can find some authors put a customized colored side bar along the top or left hand side of the heatmap to denote different groupings of the samples. You would most likely want to create a customized colored side bar based on the variable you chose to base your data reordering. The following function is based upon a code created by Dr. Peter Cock found here.
First, you will need to create a function that assigns a color to each category. This function will uses a whole bunch of “if” functions. Only the first category will be preceded by “if” and following categories will be “else if”. Add a line for each category. This example has 5 categories.
color.map<-function(diet.type){
if(diet.type=="0") "red"
else if(diet.type=="1") "green"
else if(diet.type=="2") "yellow"
else if(diet.type=="3") "blue"
else if(diet.type=="4") "purple"
}
The text in between quotation marks after “VARIABLE==” should be the exact text of the category designation within the column. So if your categories are “dry”, “muddy” and “sandy”, you should replace the numbers.
color.map<-function(CATEGORY){
if(CATEGORY=="dry") "red"
else if(CATEGORY=="muddy") "green"
else if(CATEGORY=="sandy") "yellow"
}
If the variable is continuous(such as pH), the color scheme would probably be better suited by a monochromatic scale (look at previous section for more information). Install rcolorbrewer and create a custom palette.You can change the colors and number of tones the function will produce.
install.packages("RColorBrewer")
library(RColorBrewer)
colorRampPalette(colors=c("yellow","red"))(5)
Use the hex codes outputted the last code in your color map function.
color.map<-function(CATEGORY){
if(CATEGORY=="0") "#FFFF00"
else if(CATEGORY=="1") "#FFBF00"
else if(CATEGORY=="2") "#FF7F00"
else if(CATEGORY=="3") "#FF3F00"
else if(CATEGORY=="4") "#FF0000"
}
Now that you have a function linking category and color, apply the color map to the whole column of data.
sidebarcolors <- unlist(lapply(reordermerge$VARIABLE, color.map))
Now add the argument ColSideColors=sidebarcolors to your heatmap code.
heatmap.2(justOTU2,Rowv=TRUE, Colv=FALSE, scale="column", trace="none", col=redgreen, xlab="sample", ylab="phylum", margins=c(10,15), ColSideColors=sidebarcolors)
There is actually a way to put in multiple side bars, but I have not looked much into the code. You can find an explanation here.
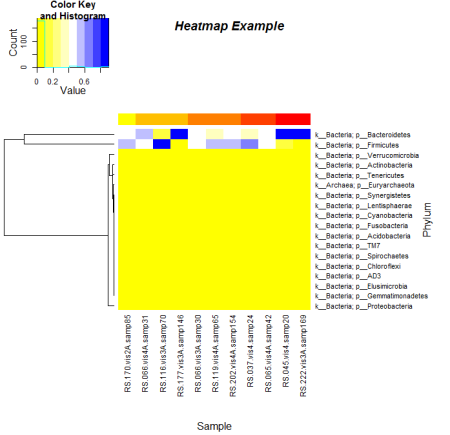
Here is an example I made with a dendrogram for rows, a customized color scale and customized sidebar colors. The color scale was created using a 9 tone palette created by rcolorbrewer.
Export the Heatmap
Once you have found the perfect way to graphically display your information as a heatmap, you will probably want to save the beauty. The export process of images is different from tables. It is similar to turning a camera on, placing an object in front of the camera, and then taking the picture. When you turn on your camera, you need to choose the size of the photo, name, and by extension, folder of the photo will be written to. The “placing the object” part is rewriting whatever final code you have come up with for your heatmap.
png(filename="output/phylum.png", width=1200, height=800) heatmap.2(justOTU2,Rowv=TRUE, Colv=FALSE, scale="column", trace="none", col=redgreen, xlab="diet type", ylab="phylum", ColSideColors=sidebarcolors, margins=c(15,15)) dev.off()
This is it! In case your curiousity about heatmaps has not been filled, there are many other aspects of heatmaps that can still be twiddled. For example, this tutorial shows how to overlay numbers on the colored boxes. If you have any suggestions to make this tutorial better, please comment below!
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.