Independent measures (between-subjects) ANOVA and displaying confidence intervals for differences in means
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
In Chapter 2 (Confidence Intervals) of Serious stats I consider the problem of displaying confidence intervals (CIs) of a set of means (which I illustrate with the simple case of two independent means). Later, in Chapter 16 (Repeated Measures ANOVA), I consider the trickier problem of displaying of two or more means from paired or repeated measures. The example in Chapter 16 uses R functions from my recent paper reviewing different methods for displaying means for repeated measures (within-subjects) ANOVA designs (Baguley, 2012b). For further details and links see a brief summary on my psychological statistics blog. The R functions included a version for independent measures (between-subject) designs, but this was a rather limited designed for comparison purposes (and not for actual use).
The independent measures case is relatively straight-forward to implement and I hadn’t originally planned to write functions for it. Since then, however, I have decided that it is worth doing. Setting up the plots can be quite fiddly and it may be useful to go over the key points for the independent case before you move on to the repeated measures case. This post therefore adapts my code for independent measures (between-subjects) designs.
The approach I propose is inspired by Goldstein and Healy (1995) – though other authors have made similar suggestions over the years (see Baguley, 2012b). Their aim was to provide a simple method for displaying a large collection of independent means (or other independent statistics). At its simplest the method reduces to plotting each statistic with error bars equal to ±1.39 standard errors of the mean. This result is a normal approximation that can be refined in various ways (e.g., by using the t distribution or by extending it to take account of correlations between conditions). Using a Goldstein-Healy plot two means are considered different with 95% confidence if their two intervals do not overlap. In other words non-overlapping CIs are (in this form of plot) approximately equivalent to a statistically significant difference between the two means with α = .05. For convenience I will refer to CIs that have this property as difference-adjusted CIs (to distinguish them from conventional CIs).
It is important to realize that conventional 95% CIs constructed around each mean won’t have this property. For independent means they are usually around 40% too wide and thus will often overlap even if the usual t test of their difference is statistically significant at p < .05. This happens because the variance of a difference is (in independent samples) equal to the sum of the variances of the individual samples. Thus the standard error of the difference is around times too large (assuming equal variances). For a more comprehensive explanation see Chapter 3 of Serious stats or Baguley (2012b).
What to plot
If you have only two means there are at least three basic options:
1) plot the individual means with conventional 95% CIs around each mean
2) plot the difference between means and a 95% CI for the difference
3) plot some form of difference-adjusted CI
Which option is best? It depends on what you are trying to do. A good place to start is with your reasons for constructing a graphical display in the first place. Graphs are not particularly good for formal inference and other options (e.g., significance tests, reporting point estimates CIs in text, likelihood ratios, Bayes factors and so forth) exist for reporting the outcome of formal hypothesis tests. Graphs are appropriate for informal inference. This includes exploratory data analysis, to aid the interpretation of complex patterns or to summarize a number of simple patterns in a single display. If the patterns are very clear, informal inference might be sufficient. In other cases it can be supplemented with formal inference.
What patterns do the three basic options above reveal? Option 1) shows the precision around individual means. This readily supports inference about the individual means (but not their difference). For example, a true population outside the 95% CI is considered implausible (and the observed mean would be different from that hypothesized value with p < .05 using a one sample t test).
Option 2) makes for a rather dull plot because it just involves a single point estimate for the difference in means and the 95% CI for the difference. If this is the only quantity of interest you’d be better off just reporting the mean and 95% CI in the text. This has advantage of being more compact and more accurate than trying to read the numbers off a graph. [This is one reason that graphs aren’t optimal for formal inference; it can be hard, for instance, to tell whether a line includes zero or excludes zero when the difference is just statistically significant or just statistically non-significant. With informal inference you shouldn’t care where p = .049 or p = .051, but whether there are any clear patterns in the data]
Option 3) shows you the individual means but calibrates the CIs so that you can tell if it is plausible that the sample means differ (using 95% confidence in the difference as a standard). Thus it seems like a good choice for graphical display if you are primarily interested in the differences between means. For formal inference it can be supplemented by reporting a hypothesis test in the text (or possibly a Figure caption).
It is worth noting that option 3) becomes even more attractive if you have more than two means to plot. It allows you to see patterns that emerge over the set of means (e.g., linear or non-linear trends or – if n per sample is similar – changes in variances) and to compare pairs of means to see whether it is plausible that they are different.
In contrast, option 2) is rather unattractive with more than two means. First, with J means there are J(J-1)/2 differences and thus an unnecessarily cluttered graphical display (e.g., with J = 5 means there are 10 Cis to plot). Second, plotting only the differences can obscure important patterns in the data (e.g., an increasing or decreasing trend in the means or variances would be difficult to identify).
Difference-adjusted CIs using the t distribution
Where only a few means are to be plotted (as is common in ANOVA) it makes sense to take a slight more accurate approach than the approximation originally proposed by Goldstein and Healy for large collections of means. This approach uses the t distribution. A similar approach is advocated by Afshartous and Preston (2010) who also provide R code for calculating multipliers for the standard errors using the t distribution (and an extension for the repeated measures). My approach is similar, but involves calculating the margin of error (half width of the error bars) directly rather than computing a multiplier to apply to the standard error.
Difference-adjusted CIs for the mean of each sample from an independent measures (between-subjects) ANOVA design is given by Equation 3.31 of Serious stats:
The term is the mean of the jth sample (where samples are labeled j = 1 to J) and
is the standard error of that sample. The
term is the quantile of the t distribution with
degrees of freedom (where
is the size of jth sample) that includes to 100(1 – α) % of the distribution.
Thus, apart from the term, this equation is identical to that for a 95% CI around the individual means, with the proviso that the standard error here is computed separately for each sample. This differs from the usual approach to plotting CIs for independent measures ANOVA design – where it is common to use a pooled standard error computed from a pooled standard deviation ( the root mean square error of the ANOVA) . While a pooled error term is sometimes appropriate, it is generally a bad idea for graphical display of the CIs because it will obscure any patterns in the variability of the samples. [Nevertheless, where
is very small it make make sense to use a pooled error term on the grounds that each sample provides an exceptionally poor estimate of its population standard deviation]
However, the most important change is the term. It creates a difference-adjusted CI by ensuring that the joint width of the margin of error around any two means is $latex \sqrt 2 $ times larger than for a single mean. The division by 2 arises merely as a consequence of dealing jointly with two error bars. Their total has to be $latex \sqrt 2 $ times larger and therefore each one needs only to be
times its conventional value (for an unadjusted CI). This is discussed in more detail by Baguley (2012a; 2012b).
This equation should perform well (e.g., providing fairly accurate coverage) as long as variances are not very unequal and the samples are approximately normal. Even when these conditions are not met, remember the aim is not to support formal inference. In addition, the approach is likely to be slightly more robust than ANOVA (at least to homogeneity of variance and unequal sample sizes). So this method is likely to be a good choice whenever ANOVA is appropriate.
R functions for independent measures (between-subjects) ANOVA designs
Two R functions for difference-adjusted CIs in independent measures ANOVA designs are provided here. The first function bsci() calculates conventional or difference-adjusted CIs for a one-way ANOVA design.
bsci <- function(data.frame, group.var=1, dv.var=2, difference=FALSE, pooled.error=FALSE, conf.level=0.95) {
data <- subset(data.frame, select=c(group.var, dv.var))
fact <- factor(data[[1]])
dv <- data[[2]]
J <- nlevels(fact)
N <- length(dv)
ci.mat <- matrix(,J,3, dimnames=list(levels(fact), c('lower', 'mean', 'upper')))
ci.mat[,2] <- tapply(dv, fact, mean)
n.per.group <- tapply(dv, fact, length)
if(difference==TRUE) diff.factor= 2^0.5/2 else diff.factor=1
if(pooled.error==TRUE) {
for(i in 1:J) {
moe <- summary(lm(dv ~ 0 + fact))$sigma/(n.per.group[[i]])^0.5 * qt(1-(1-conf.level)/2,N-J) * diff.factor
ci.mat[i,1] <- ci.mat[i,2] - moe
ci.mat[i,3] <- ci.mat[i,2] + moe
}
}
if(pooled.error==FALSE) {
for(i in 1:J) {
group.dat <- subset(data, data[1]==levels(fact)[i])[[2]]
moe <- sd(group.dat)/sqrt(n.per.group[[i]]) * qt(1-(1-conf.level)/2,n.per.group[[i]]-1) * diff.factor
ci.mat[i,1] <- ci.mat[i,2] - moe
ci.mat[i,3] <- ci.mat[i,2] + moe
}
}
ci.mat
}
plot.bsci <- function(data.frame, group.var=1, dv.var=2, difference=TRUE, pooled.error=FALSE, conf.level=0.95, xlab=NULL, ylab=NULL, level.labels=NULL, main=NULL, pch=21, ylim=c(min.y, max.y), line.width=c(1.5, 0), grid=TRUE) {
data <- subset(data.frame, select=c(group.var, dv.var))
if(missing(level.labels)) level.labels <- levels(data[[1]])
if (is.factor(data[[1]])==FALSE) data[[1]] <- factor(data[[1]])
if (is.factor(data[[1]])==TRUE) data[[1]] <- factor(data[[1]])
dv <- data[[2]]
J <- nlevels(data[[1]])
ci.mat <- bsci(data.frame=data.frame, group.var=group.var, dv.var=dv.var, difference=difference, pooled.error=pooled.error, conf.level=conf.level)
moe.y <- max(ci.mat) - min(ci.mat)
min.y <- min(ci.mat) - moe.y/3
max.y <- max(ci.mat) + moe.y/3
if (missing(xlab))
xlab <- "Groups"
if (missing(ylab))
ylab <- "Confidence interval for mean"
plot(0, 0, ylim = ylim, xaxt = "n", xlim = c(0.7, J + 0.3), xlab = xlab,
ylab = ylab, main = main)
grid()
points(ci.mat[,2], pch = pch, bg = "black")
index <- 1:J
segments(index, ci.mat[, 1], index, ci.mat[, 3], lwd = line.width[1])
segments(index - 0.02, ci.mat[, 1], index + 0.02, ci.mat[, 1], lwd = line.width[2])
segments(index - 0.02, ci.mat[, 3], index + 0.02, ci.mat[, 3], lwd = line.width[2])
axis(1, index, labels=level.labels)
}
The default is difference=FALSE (on the basis that these are the CIs most likely to be reported in text or tables). The second function plot.bsci() uses the former function to plot the means and CIs the default here is difference=TRUE (on the basis that it the difference-adjusted CIs are likely to be more useful for graphical display). For both functions the default is a pooled error term (pooled.error=FALSE) and a 95% confidence level (conf.level=0.95). Each function also takes input as a data frame and assumes that the grouping variable is the first column and the dependent variable the second column. If the appropriate variables are in different columns, the correct columns can be specified with the arguments group.var and dv.var. The plotting function also takes some standard graphical parameters (e.g., for labels and so forth).
The following examples use the diagram data set from Serious stats. The first line loads the data set (if you have a live internet connection). The second line generated the difference-adjusted CIs. The third line plots the difference adjusted CIs. Note that the grouping variable (factor) is in the second column and the DV is in the fourth column.
diag.dat <- read.csv('http://www2.ntupsychology.net/seriousstats/diagram.csv')
bsci(diag.dat, group.var=2, dv.var=4, difference=TRUE)
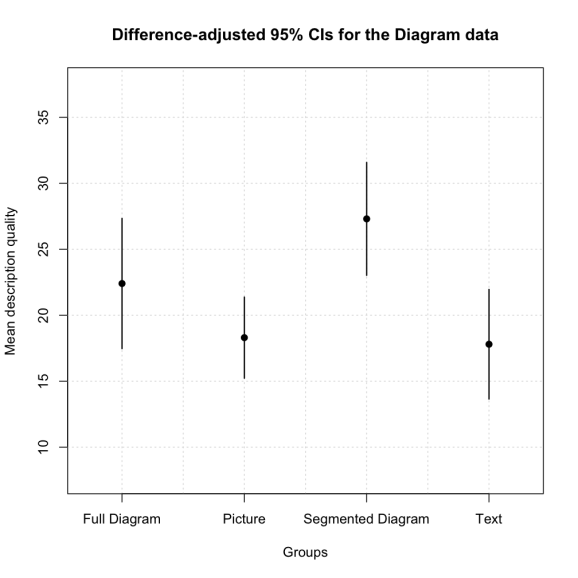
plot.bsci(diag.dat, group.var=2, dv.var=4, ylab='Mean description quality', main = 'Difference-adjusted 95% CIs for the Diagram data')
In this case the graph looks like this:
It should be immediately clear that while the segmented diagram condition (S) tends to have higher scores than the text (T) or picture (P) conditions, but the full diagram (F) condition is somewhere in between. This matches the uncorrected pairwise comparisons where S > P = T, S = F, and F = P = T.
At some point I will also add a function to plot two-tiered error bars (combining option 1 and 3). For details of the extension to repeated measures designs see Baguley (2012b). The code and date sets are available here.
References
Afshartous D., & Preston R. A. (2010). Confidence intervals for dependent data: equating nonoverlap with statistical significance. Computational Statistics and Data Analysis. 54, 2296-2305.
Baguley, T. (2012a, in press). Serious stats: A guide to advanced statistics for the behavioral sciences. Basingstoke: Palgrave.
Baguley, T. (2012b). Calculating and graphing within-subject confidence intervals for ANOVA. Behavior Research Methods, 44, 158-175.
Goldstein, H., & Healy, M. J. R. (1995). Journal of the Royal Statistical Society. Series A (Statistics in Society), 158, 175-177.
Schenker, N., & Gentleman, J. F. (2001). On judging the significance of differences by examining the overlap between confidence intervals. The American Statistician, 55, 182-186.
N.B. R code formatted via Pretty R at inside-R.org
Filed under: R code, serious stats, stats advice Tagged: ANOVA, confidence intervals, exploratory data analysis, psychology, R, repeated measures ANOVA, significance tests, statistics
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.