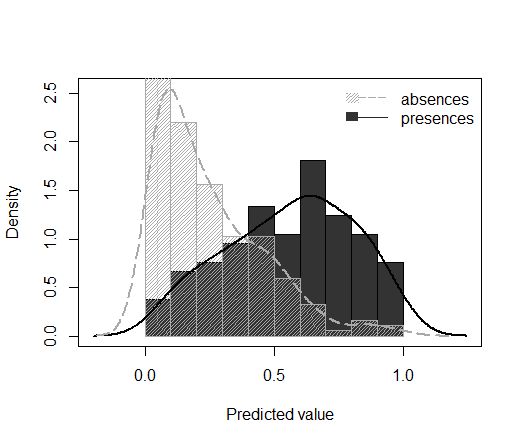
Plot the density of predicted values for presences and absences
[This article was first published on modTools, and kindly contributed to R-bloggers]. (You can report issue about the content on this page here)
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
The ‘predDensity’ function, included in the modEvA package since version 1.5 (currently available on R-Forge), produces a histogram and/or a kernel density plot of predicted values for observed presences and absences in a binomial GLM:
predDensity <- function (model = NULL, obs = NULL, pred = NULL, separate = TRUE,
type = c("both"), legend.pos = "topright")
{
if (!is.null(model)) {
if (!("glm" %in% class(model)) || family(model)$family !=
"binomial")
stop("'model' must be of class 'glm' and family 'binomial'.")
if (!is.null(obs))
message("Argument 'obs' ignored in favour of 'model'.")
if (!is.null(pred))
message("Argument 'pred' ignored in favour of 'model'.")
obs <- model$y
pred <- model$fitted.values
}
if (is.null(obs)) {
if (is.null(pred))
stop("You must provide either 'model' or 'pred'.")
separate <- FALSE
obs <- sample(c(0, 1), length(pred), replace = TRUE)
}
else {
if (length(obs) != length(pred))
stop("'obs' and 'pred' must have the same length.")
}
pred0 <- pred[obs == 0]
pred1 <- pred[obs == 1]
type <- match.arg(type, c("histogram", "density", "both"))
rslt <- vector("list")
if (type %in% c("density", "both")) {
if (!separate) {
dens <- density(pred)
xrange <- range(dens$x, finite = TRUE)
yrange <- range(dens$y, finite = TRUE)
rslt[["density"]] <- dens
}
else {
dens0 <- density(pred0)
dens1 <- density(pred1)
xrange <- range(dens0$x, dens1$x, finite = TRUE)
yrange <- range(dens0$y, dens1$y, finite = TRUE)
rslt[["density_obs1"]] <- dens1
rslt[["density_obs0"]] <- dens0
}
plot(x = xrange, y = yrange, xlab = "Predicted value",
ylab = "Density", type = "n")
}
if (type %in% c("histogram", "both")) {
hist0 <- hist(pred0, plot = FALSE)
hist1 <- hist(pred1, plot = FALSE)
if (type == "histogram") {
yrange <- range(hist0$density, hist1$density, finite = TRUE)
plot(x = c(0, 1), y = yrange, type = "n", xlab = "Predicted value",
ylab = "Density")
}
if (!separate) {
histogram <- hist(c(pred0, pred1), freq = FALSE,
col = "grey20", add = TRUE)
rslt[["histogram"]] <- histogram
}
else {
hist(pred1, freq = FALSE, col = "grey20", add = TRUE)
hist(pred0, freq = FALSE, col = "darkgrey", density = 40,
angle = 45, add = TRUE)
rslt[["histogram_obs1"]] <- hist1
rslt[["histogram_obs0"]] <- hist0
if (legend.pos != "n" && type == "histogram")
legend(legend.pos, legend = c("absences", "presences"),
fill = c("darkgrey", "grey20"), border = NA,
density = c(40, NA), bty = "n")
}
}
if (type %in% c("density", "both")) {
if (!separate) {
lines(dens, col = "black", lwd = 2)
}
else {
lines(dens1, col = "black", lwd = 2)
lines(dens0, col = "darkgrey", lty = 5, lwd = 2)
if (legend.pos != "n" && type == "density")
legend(legend.pos, legend = c("absences", "presences"),
col = c("darkgrey", "black"), lty = c(5, 1),
bty = "n")
if (legend.pos != "n" && type == "both")
legend(legend.pos, legend = c("absences", "presences"),
fill = c("darkgrey", "grey20"), border = NA,
lty = c(5, 1), col = c("darkgrey", "grey15"),
density = c(40, NA), bty = "n")
}
}
return(rslt)
}
Usage example:
install.packages("modEvA", repos="http://R-Forge.R-project.org")
library(modEvA)
data(rotif.mods)
predDensity(model = rotif.mods$models[[3]])
To leave a comment for the author, please follow the link and comment on their blog: modTools.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
