Data Chats: An Interview on Data-Driven Campaigns, Bias & Ethics
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
Background
One of the motives for starting the Data Chats interview series was to shed light on the many ways in which data and analytics professionals operate across different fields and cultures. Previously, Avision Ho (Senior Data Scientist at the British Department for Education at the time) and Abhishek Modi (Data Science Consultant at Deloitte at the time) described the data science career journey and answered technology-specific questions (e.g. favourite R packages). So I thought I’d do an interview on how analytics is applied in a very different, yet important, setting: politics.
This time, I have the pleasure to speak with Christopher Treshan Perera and Nanthida Rakwong from Worldacquire, a digital consultancy with politics as a core practice area. They launched with a mission to use analytics and digital tech in political campaigns, public affairs and human rights. They notably managed campaigns at the 2019 Thailand general election and the 2019 Hong Kong District Council elections; at the latter, they helped a pro-democracy candidate defeat a long-standing incumbent. Their co-founders have spoken at the United Nations and the UK Parliament on ethical issues in technology and society1 and managed research for Konrad-Adenauer-Stiftung, Germany’s governing party think tank.
In this in-depth interview, Christopher and Nanthida discuss how they navigate analytics and politics, challenges they encountered (e.g. how to obtain reliable data in Thailand), ethical questions (Cambridge Analytica, GDPR) and other practical considerations.
The Interview
M: It’s great to have you guys here. Tell me a bit about Worldacquire and your journey to bringing together analytics and politics!
C: We launched two years ago to explore how AI and data-driven technology could enhance democracy – whether by helping aspiring politicians win elections or by supporting research and campaigning efforts of parties and public organisations.
N: I grew up in Thailand, but moved to London in 2010 to work as a political consultant for Amsterdam and Partners, an international law firm representing political figures. My most important case was in Thai politics – bringing the Thai junta (military government) to justice at the International Criminal Court (ICC) over the 2010 Bangkok Massacres. I also advised the “Red Shirts” pro-democracy movement in Thailand and various other political parties around the world.

C: I have a more corporate and techie background. My career started as a data analyst at Bloomberg in London, followed by several years at viagogo, an online marketplace for sports and show tickets, where I rose from digital marketing exec to global advertising management. It is common to wear many hats at a tech startup and mine included data analysis, business intelligence, product management and algorithm design for marketing APIs. Then I moved to American Express where my task was to transfer digital marketing know-how from the tech world to “big finance”.
Before Worldacquire I already wanted to connect tech with social causes. Back in 2015 I founded Outreach Digital, an entirely volunteer-run association making digital skills more accessible; it has since become the largest meetup group in London’s tech and digital space.
M: That’s a great fusion of politics and digital marketing. Also, Chris – we first met at one of those meetup groups, which makes a great case for their networking value. So what exactly got you thinking: wow, it would be great to merge analytics and political consulting?
N: While I was advising the Thai pro-democracy movement, I realized how crucial it was to understand a situation in real-time in order to make better decisions. By collecting a lot of data, compiling and analysing it faster, we could operate in a more efficient and scalable way. That led to my interest in data, analytical technologies and AI, and I immediately saw their dangers, too: the Cambridge Analytica scandal, for example, was a misuse of those technologies. But you can change something for the better only if you engage and participate in shaping it.
M: Using AI and data to make better decisions – could you describe how you can do that in politics? How do other organizations do that today?
N: Let’s say you want to understand millions of people including your potential voters. By gathering data from multiple sources, including Facebook, Twitter, forums, emails and more, you can find new paths to make everyone work together towards the same goal. If you want to run a political campaign of any kind but don’t understand what exactly people want, it’s very hard to bring them together.
Moreover, you always need to identify new supporters. There may be people who are unsure about your cause or movement – perhaps they are friends of your ardent supporters – but they hesitate to join because they don’t understand it well enough. You can use AI and data technology to understand what they need and how to best communicate with them.
C: One of the things Cambridge Analytica did was to uncover a blind spot in the political “market share” or electorate and target those swing voters (and people who never wanted to vote) using unethical advertising methods, including fake news.
This raises the question of whether the technology can be used in an ethical and transparent way. If everyone was aware of the practices, if parties or candidates communicated transparently about them, then perhaps we’d have a different situation even in the UK right now. The other consideration is which specific AI and analytics methods to apply: you could implement recommendation systems, pattern recognition techniques or combine existing methods.
N: And use them to spread truth and support good causes, rather than rumours, disinformation and division.
M: So you do the opposite of what Cambridge Analytica did by using the technologies in a more ethical and transparent way?
N: That’s right. If we do not engage in this game, we basically allow malicious or unethical players to misuse it. If we withdraw we cannot help shape new solutions and perspectives.
C: Think practically! Advertising has a very long history – newspaper, TV, radio and billboards. Over the decades every new medium got increasingly regulated, but advertising still exists and it has arguably become more transparent and ethical. If traditional advertising channels can improve over time, so can digital advertising.
M: You also believe that by using these techniques for a good cause, we can make our democratic systems more resilient and less prone to being abused?
C&N: Correct! Moreover, they can also be used to improve public services and government-to-citizen communications, especially during a crisis.
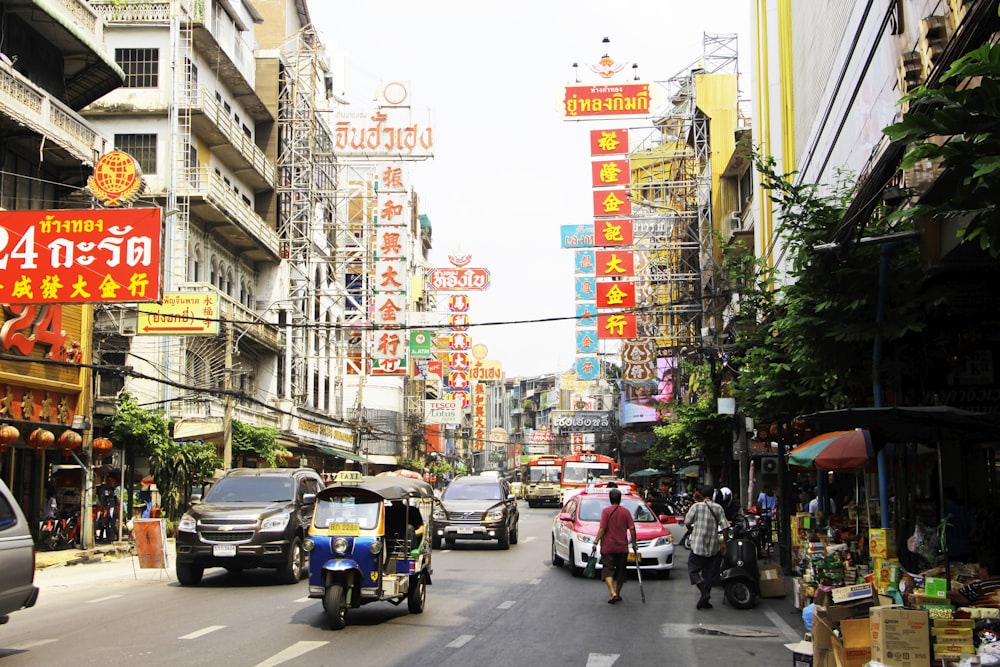
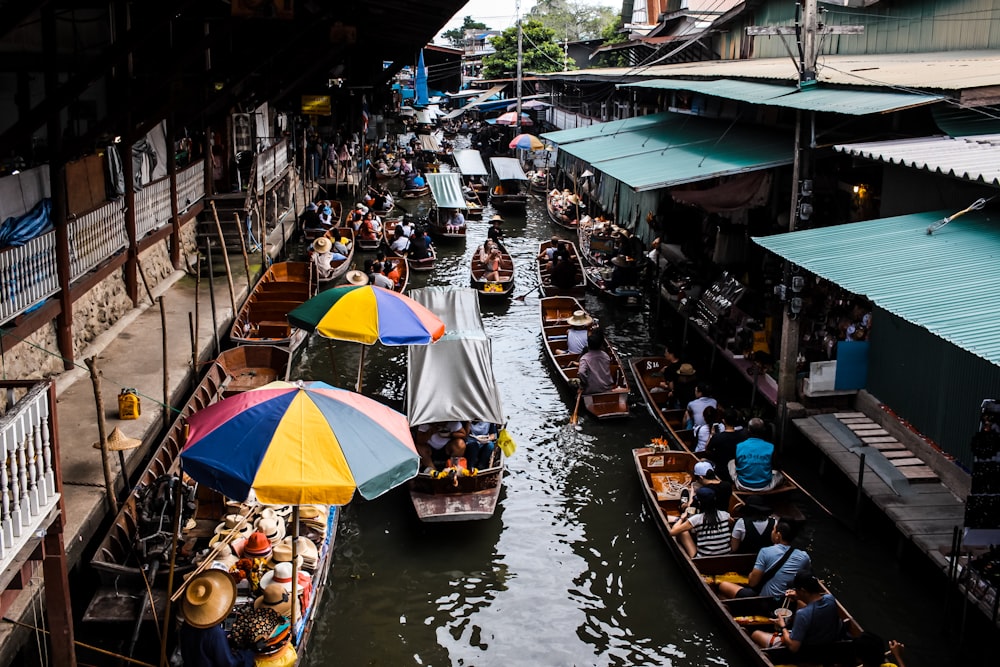
M: Now let’s talk about the election campaign that you guys managed in Thailand. Could you tell me more about it?
N: First of all, it was a long-awaited election because Thailand has been ruled by an authoritarian military junta since 2014. We advised a first-time candidate from a new party standing for MP in Bangkok. Candidates were given only six weeks to campaign – a major challenge considering this was one of the largest constituencies.
Without any data to start with, we went to the local administration office to request the electoral register, but they essentially refused to share anything. We suspected this happened because of the deep influence of the incumbent. We had to think differently!
We could have started canvassing (surveying and campaigning) from door to door, but this would have been an issue for several reasons: firstly, Thai people don’t vote based on where they reside, but where their home was registered. Thus, people who live in a constituency may not have the right to vote there. Secondly, the dangerous political climate and Thailand’s harsh censorship laws made people extremely wary of sharing their political views or past voting behaviour.
Another option was to collect data and communicate online through social media advertising. Unfortunately the party leadership wasn’t willing to invest in it. They preferred to play it safe and spend money on leaflets and billboards instead.
We ultimately went for a sampling method using fieldwork mapping: we divided the constituency into smaller areas based on the polling stations that cover them and interviewed a sample of people in each area (this was still challenging considering how wary people were!) and built our understanding of the overall constituency based on data. We facilitated this by using an app called Mela.


C: We didn’t use any advanced AI magic in this instance, but the project highlighted how important it is to get the right data and to ensure that the initial dataset is clean. What data scientists often do is go on the internet and pick whatever datasets are available online – but these are often outdated, incomplete or even biased.
Especially in a developing country with poor accountability and no balance of powers, it is hard to verify if research and survey data is correct. You have to go hands-on and create the conditions for people to share their genuine views. Once the data is accurate, you can start doing advanced stuff.
N: We would have certainly received more valuable insights had the party leadership approved social media advertising. It takes more effort to measure the conversion rate of leaflets and billboards. Especially if you only have six weeks to campaign.
C: Nonetheless we were able to run a smaller-scale test and get accurate social insights. Ideally, we would have gathered enough data to run prediction and network algorithms to evaluate the profile, behaviour and preferences of each constituency sub-area, and thereby understand which political issues mattered to them the most. Then we could have tailored the messages that would best resonate with each group. Despite the small amount of data, it was enough to draw some important conclusions about the potential voters.
M: It really makes sense when facing such time constraints, so it’s a shame that this is underutilized in election campaigns. Chris, what is the biggest difference between digital advertising in a commercial vs a political setting?
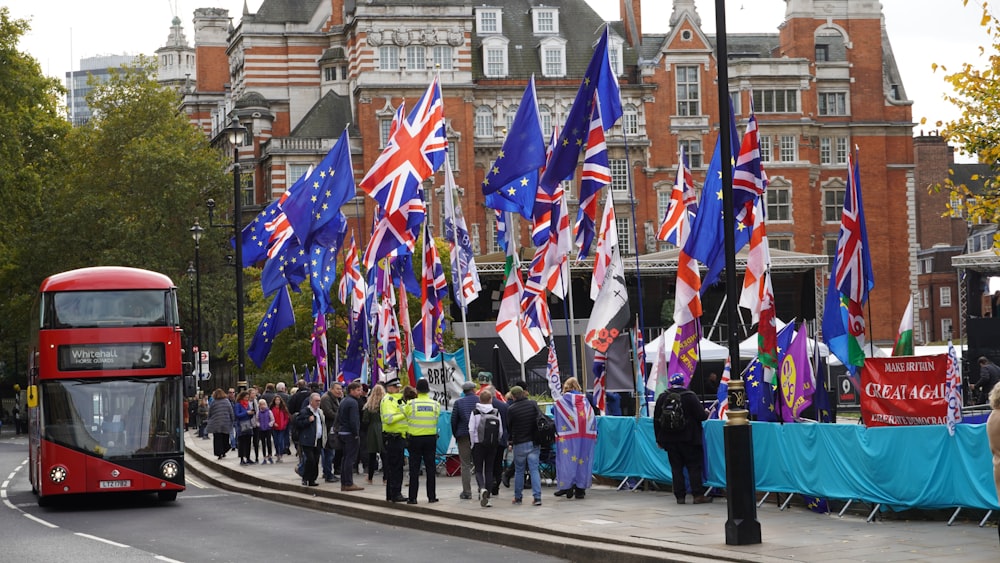
C: In business you have more wiggle space for your message choice. You can use superlatives like “best tickets” or “best concerts”. In politics people do that, too, but it can be dangerous; think about the Brexit bus with the exaggerated claim about the NHS money. There will also be very different budgets due to both internal and external factors. Businesses are often more willing to invest in advertising as it leads to direct and immediate sales. In politics, the “sales cycle” can take much longer.
N: In many countries, including Thailand, there’s also a legal cap on campaigning spend.
C: Another issue is visibility. Most digital platforms decide which content to display based on ranking algorithms; one factor that influences those algorithms is pre-existing activity and performance. For example, if you want to advertise a car on Google, and you build a Google Ads campaigns around the keyword “car”, Google’s algorithms will already know that this is something businesses want to advertise based on historic performance data. The algorithms will also know that people click on those ads after searching “car”. On the contrary, if a keyword was never used before or isn’t typically associated with people clicking on ads, Google will wait a little longer before displaying ads for it. So there can be some delays before an ad for a new politician is actually visible, but it is still faster than trying to get a blog post go viral.
M: So there’s both a legal and a search engine strategy aspect. How about analytics in business vs politics?
C: There are many similarities. You just need to translate a concept from one field into another. In business, KPIs and metrics are formed around impressions, actions (sales), CTR and conversion rates. In politics it’s more about long-term performance, maybe along the lines of CLV (customer lifetime value).
N: Having many clicks on your ad or post doesn’t necessarily translate into votes. It could even be negative – think about Prince Andrew!
M: What were your top challenges at the Thai election?
N: The lack of accurate data and the poor awareness about the importance of data by the leadership – especially current or real-time data. People still rely on old reports and outdated information.
M: What was the outcome of the election campaign?
N: Our candidate lost, but exceeded our expectations. More importantly, the winning candidate was from the Future Forward Party, another new and allied party that did invest significantly in its social media at a national level. We observed that they also used tailored, targeted advertising and A/B-testing to gather data about voter preferences. The political party’s image really helped that candidate. Like our candidate, he was not a resident of the constituency yet still won. This was the very first time that anyone used social media as a key channel for data collection in an election campaign in Thailand.
C: Indeed, this was forward-thinking. Many political campaigners around the world use social media, but don’t make the most of its advanced algorithms and data-gathering capabilities. Considering the difficulties we had in accessing data, one of the biggest learnings is that even in the face of authoritarian red tape and bureaucracy, digital platforms can help overcome hurdles in understanding your audience.
M: Let’s talk about regulation and ethics. Is data is the new oil? A valuable resource that helps society progress?
C: Yes, it can be – depending on which society it will be used in. If a society has strong data protection and privacy laws, then data can be used hand-in-hand with democratic principles. If not, then it can be a very “bad oil”.
Looking back in time, radio was “the new oil” at some point. TV, too. From a regulatory point of view, they all provoked concerns (including about propaganda), but over time different bodies and regulations were formed to address them, such as today’s ASA in the UK; and now for data and digital technologies, we have the ICO in the UK.
M: What is your position regarding GDPR? Also, when people think about applying analytics in political advertising, many worry about Cambridge Analytica and Brexit. Obviously you want to promote good causes and democracy – but can you avoid repeating their mistakes?
N: Let’s start with Obama’s presidential campaign. Obama used similar tactics, but what did he do differently? He informed people about how their data would be used. So transparency is really a key factor.
C: I think there can be some misunderstanding that influences the public’s perception, but also that of advertisers. Some advertisers (including businesses and politicians) can see GDPR as a nuisance, when in fact it is an opportunity. In the same way that media regulation helped build trust in radio and TV, GDPR and future internet or data protection laws will increase trust in digital spaces. Once people feel safe about the new media, they will feel even more comfortable using the platforms and sharing their data.
When people know that there is a rule of law and accountability around data, they will feel less suspicious than with the dodgy websites today that say very little about data protection. At the same time, GDPR will push companies – including advertisers – to design their products, solutions and activities around privacy. This should help encourage ethical uses of this “new oil”.
As a matter of fact, there will be an increasing number of new regulations that will touch upon different issues in the realm of data-driven products, notably in AI. Algorithmic bias, for instance, is one hot topic at the moment. Since AI is based on algorithms that learn from historical data, that data could reinforce existing social biases – whether it’s about predicting who will be the next criminal, or who deserves a visa or insurance. How to solve this? Regulation can and needs to answer this, whether by requiring AI products to make their program code public or by creating mechanisms to prevent or defeat the bias.
N: People need to understand how software works. Once again, transparency is vital. Many of the issues around algorithms and the misperceptions of AI being dangerous come down to the fact that the technical issues are not properly explained. When AI experts and advocates come together and make an effort to ensure that biases and manipulation are eliminated from these algorithms, that’s when products and solutions based on AI can be more ethical and transparent.
M: It’s important to understand where all the data people provide goes and I believe there’s also the question of consent, which is covered by GDPR. What I find really interesting is algorithmic bias, but also the idea that people can better judge what is right and wrong only after they are educated about how the algorithm works.
C: Exactly, and this is what regulators are starting – and should be starting – to think about.
There are some who think that political advertising should be completely banned. Twitter went down this road and many applauded their decision, with the main perception being that data and algorithms can be misused the way Cambridge Analytica did. However, what was ignored was the fact that all these algorithms don’t need to work in a black box – in fact, they can be revealed, changed, overridden.
For example, a recommendation system could, instead of saying “A is better than B”, explain “We recommend A over B because our algorithm observed that you like x, y and z.” Companies may be reluctant to fully reveal their algorithm code, but they could at least give an idea of what parameters are taken into account, what outcomes can be expected, and why. Once again, transparency is key.
Another aspect ignored in the whole debate about political advertising is the fact that if digital platforms like Twitter ban them, newcomer politicians will struggle to gain a following or communicate with their target audience if they have a time constraint, like in the case of Thailand. Banning political ads carries huge disadvantages, especially for politicians who already lack resources.
The right approach would have been to push for more transparency – not only in advertising (“paid”) algorithms, but also in the “organic” and “earned” algorithms used on the very same social media platforms.
N: Moreover, Twitter appears not to care as much as they say about what content is posted on the platform. If they can ban political ads, why do they do so little against hate speech and other online harms? Many, including myself, have experienced harassment and public death threats on Twitter, yet Twitter refused to cooperate swiftly and proactively with the British police – instead they shifted the burden of proof to the victims.
M: It is also very unclear how they will actually implement the ban.
N: I honestly think it’s dangerous that Jack Dorsey (the CEO of Twitter) calls for “earned” popularity and encourages a culture of going viral. If a tweet goes viral does it really mean that it “earned” it? Is it really more accurate and correct than other tweets? More than a few times, a viral message has spread false accusations and fake news.
C: To make things worse, going viral is also determined by algorithms. How does Twitter decide which content should get more visibility? Is it the likes, the retweets, the popularity of the tweet author and their followers? We have seen (and tested) how easily this can be manipulated.
So banning political ads doesn’t solve the problem of powerful obscure algorithms, as organic content is decided by even less transparent ones! Typically, such algorithms seem to favour users who already have a strong following – in the case of an election, this is often the incumbent. Newcomers can be heavily penalized by this dynamic.
Another issue are fake users and bots on Twitter. These can be bought in thousands or more to mass-like or mass-follow and exaggerate the popularity of a particular user or a post. Equally, your competitor could get 10,000 Twitter bots or fake users to report your public posts as spam or abusive. This is an easy way to manipulate the system (also tested in Thailand). The Twitter algorithm will likely disregard the fact that the users are fake and make the falsely reported tweet disappear even if it is genuine and popular.
M: This is something I have seen in Hong Kong, too. Do you think censorship is a solution?
C: Did TVs ban political content? Do billboards ban political ads? Not really, so we really don’t think that banning political ads is a solution.
N: It should be more about regulating content, what is not OK to say.
M: How do you differentiate yourselves from other consultancies or agencies that do the same or similar things as you?
N: Firstly, aside from working on the big picture strategy we actually also implement it. Working hands-on gives us a much more tangible picture of the dynamics, limitations and issues that could be faced.
C: When you hear academics, researchers and thought leaders jump enthusiastically to praise Jack Dorsey for banning political ads on Twitter, a practitioner who personally set up digital campaigns for politics will tell you how disastrous the effects of his decision can be. Our USP is that we work on the ground and thoroughly understand technical implications and their consequences.
M: This ties back with what you mentioned about ‘blind spots’. Being on the ground helps you spot them, too, right?
N: Correct, and it also helps with finding alternative solutions in case Plan A didn’t work in the first place.
M: What is your vision for your business?
C: We truly believe that AI, data and digital technologies can be used for good causes – and we want to show that this is true and applies to anywhere around the world. It is also important for people to understand the uses of these technologies and the actors who control them. We want to help people understand both sides of the coin. Many governments and NGOs have digital and data on their agendas, but often seem to have a very superficial sense of the technologies – we want to help there. And we want to be involved in and lead the societal, political and ethical debates around these technologies, as well as demystify the exaggerated perceptions of danger.

M: Is there any advice you have for data scientists who aspire to work on political projects, political data scientists, or data scientists in general?
C: Get the right data, know the sources and eliminate biases! This includes statistical bias (especially sampling, funding and reporting bias), cognitive bias and other social biases. Online datasets may be easier to obtain, but are outdated or not real time enough. Worse, they could be doctored to reflect the narratives of an authoritarian regime or lobbyist group.
Ask yourselves: what were the context and conditions during the data collection process? What kind of limitations existed? Whether it’s data from a sentiment analysis report or a simple survey, what could be wrong with the data? Could there be any noise? Anything unusual?
Also, the logic behind the metrics in a dataset can be misleading; think about GDP, a measure for economic growth. Does a growing GDP mean the country is improving and everyone is better off? Not really. If you look closer you might see that the GDP growth is distributed only to a small percentage of the population.
How were survey responses recorded? Did the method change during the campaign? What’s the logic behind the metrics? What kind of issues may lead to the data being wrong? Could there be a situation of reluctant journalists or silenced human rights activists?
Finally, ethics is not only for philosophers, but also for engineers. This will be a hot topic over the next years, and AI and data specialists will need to be able to explain to consumers and other stakeholders the different problems and solutions in algorithm-driven products.
N: In politics and economics especially, it is really important to ask who created the dataset, who financed or sponsored it, and who really controls the overall character and narrative of the data.
C: You should never be afraid to go out there on the field and collect the data by yourself – it can be really fun!
M: Thank you again for your time and for this very fascinating interview.
Endnotes
What I thought was an interesting theme is the ubiquity in the application of analytics. But some of the data challenges that Nanthida and Chris raised are very real, and confirms the view that a considerable chunk of time in data analysis is spent on collecting, cleaning and getting the data right for analysis in the first place, not only the analysis itself.
I hope you’ve enjoyed reading the above interview. If you would like to get in touch with Christopher and Nanthida, you may reach them through their website here. I’m also looking to do more interviews, so if you are a data / analytics practitioner and you think you have something interesting to share, please feel free to get in touch!
Image credits: Artem Bryzgalov, Kelvin Yup, Frida Aguilar Estrada, Franz Wender, ThisisEngineering, camilo jimenez, dole777, jbdodane, History in HD, Jonathan Francisca, Carlos Muza
See UN-related work on digital ethics and transparency, and on anti-disinformation: https://worldacquire.com/2019/12/09/worldacquire-at-the-united-nations-igf-2019/ and https://worldacquire.com/2020/02/27/online-disinformation-and-extremism-how-it-spreads-and-how-to-stop-it/↩︎
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.